Group:SMART:Tangible Models of Cdc42 Interacting With Intersectin
From Proteopedia
Contents |
A Lincoln HS SMART Team Telling a Molecular Story
Abraham Lincoln High School, San Francisco, California
- Students -- Cyndi He, Glen Huynh, Aissa Isana, Bradley Jann, Bradford Li, Yaqiao Li, Calvin Ng, Tiffany Saw, Elizaveta Sergeeva, Jacqueline Tam, Allison Trinh, Michelle Xie, Ellen Zhang
- Teacher -- Julie Reis
Mentors:
- UCSF Science and Health Education Partnership -- Sabine Jeske
- UCSF/UCB Joint Graduate Group in Bioengineering -- Angela Chau
- Institute for Neurodegenerative Diseases, UCSF -- Kurt Giles
- Center for BioMolecular Modeling, Milwaukee School of Engineering -- Shannondoah Colton
Abstract for Our Project
Abstract
- Information transfer within the cell relies upon signaling pathways made up of interacting proteins. Rho-family GTPases are GTP-binding proteins, which function as molecular switches initially in the "off" state. Interactions with activators turn these GTPases "on" and they then interact with other proteins, leading to a variety of cellular behaviors such as directed movement and changes in cell shape.
- Traditional methods for visualizing protein structures and interactions are often limited in the amount of information that can be conveyed in two dimensions. Using the rapid prototyping technology at the Milwaukee School of Engineering's Center for BioMolecular Modeling, we have built tangible 3D models of the Rho GTPase Cdc42 in complex with one of its activators. Our model shows the interaction of Cdc42 with the catalytic domains of intersectin, based upon the published crystal structure solved by Snyder, et al (PDB 1KI1, Nature Structural Biology 2002; 9(6): 468- 475). Along with computer visualization tools, tangible 3D models allow students and scientists alike to more fully explore the intricacies of protein interactions.
Cell Signaling and Cell Movement
- Cell signaling is how cells transmit information from the outside environment to inside the cell as well as how cells propagate messages within the cell. Transmembrane receptors at the cell surface detect environmental changes and pass the information on to intracellular signaling proteins. Proteins that make up signaling pathways pass along messages by interacting with each other.
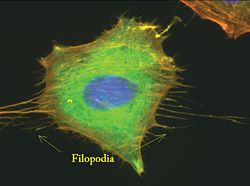
- Cell movement is integral for survival in both bacterial and eukaryotic cells. In eukaryotes, cells move by changing their cytoskeletons to form actin structures like filopodia. These structures are formed in response to signals from outside of cells which are relayed throughout the cell using signaling pathways.
Why build a model?
- A 3-dimensional model generated by the polymer printing technology at the Center for BioMolecular Modeling greatly enhances our ability to visualize and explore Cdc42 and its interaction with Intersectin. The complex structures and shapes of these molecules can be appreciated more fully through seeing and touching a physical model. A tangible model can not only aid us in summarizing what we know but can also inspire us and other scientists to ask new questions and come up with new hypotheses about these molecules.
Cdc42: The GTPase
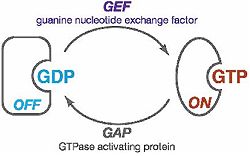
- Like other Rho GTPases, can exist in either the “Off” or “On” state. A guanosine diphosphate (GDP) in the nucleotide-binding pocket indicates the “Off” state, and a guanosine triphosphate (GTP) indicates the “On” state. The activation from the “Off” state to the “On” state is catalyzed by a guanine nucleotide exchange factor (GEF). Thee hydrolysis of GTP, which turns the GTPase off, is catalyzed by a GTPase activating protein called a GAP.
- In addition to binding to a nucleotide (GDP or GTP), Cdc42 is usually found to be bound to a magnesium ion (Mg++), shown in magenta. The Mg++ sits near the nucleotide-binding pocket and helps to stabilize the Cdc42-nucleotide interaction.
Intersectin - The GEF
- Intersectin is one of the activators of the GTPase Cdc42. It is a guanine nucleotide exchange factor (GEF) and turns on Cdc42 by catalyzing the exchange of GDP for GTP. Intersectin is a signaling protein that contains many domains, not all of which are shown here. The structure of the Cdc42/Intersectin complex was solved with the DH (residues 1229- 1439) and PH (residues 1440-1580) domains of Intersectin. Here, we focus only on the DH domain because the PH domain does not appear to come into any significant contact with Cdc42.
How Intersectin Activates Cdc42
- The orientation of Alanine 59 is the highlight of Cdc42 activation by Intersectin. Alanine 59 causes the displacement of the Mg++ ion which in turn releases GDP from Cdc42’s binding pocket. Cdc42 is then able to accept a new nucleotide, specifically GTP, to become activated.
|
1) ( yellow ) is off when GDP is in its binding pocket. Mg++ is shown in magenta, Alanine 59 in green, Switch 1 in light gold, and Switch 2 in dark orange. Brown depicts the residues interacting with GDP (Lys16 and Cys18).
2) (blue) binds to Cdc42, and its DH domain interacts with the GTPase.
3) Thee interaction causes conformational changes in Switch 1 and Switch 2 of Cdc42. Thr35 hydrogen bonds to a residue (Glu1244) of the DH domain of Intersectin. Most important is the noticeable change in the orientation of residue 59 (Ala59). Its sidechain flips into the magnesium-binding pocket, causing the release of the Mg++ ion and in turn the release of GDP from Cdc42.
4) Mg++ and GDP fall off from Cdc42. The nucleotide-binding pocket of Cdc42 is now empty.
5) Because the intracellular ratio of GTP to GDP is high, a GTP molecule (dark green) will float into the binding pocket of Cdc42 and activate it.
Animation: How Intersectin Activates Cdc42
MSOE Center for BioMolecular Modeling and SMART Teams
SMART Teams (Students Modeling A Research Topic) is a science outreach program developed by the MSOE Center for BioMolecular Modeling. In this program, teams of high school students work with a local resarch lab to design and build a physical model of a protein that is being investigated by the lab. The goal of the SMART Team program is to introduce students to the real world of scince --- as it exists in a local research lab. The development of this program was supported by grants from the NIH-NCRR SEPA program (Science Education Partnership Award) and an HHMI Precollege Science Education Award. For more information about this program, visit the CBM web site at www.rpc.msoe.edu/cbm .