Molecular Playground/HypA
From Proteopedia
References
- ↑ Kuipers EJ. Helicobacter pylori and the risk and management of associated diseases: gastritis, ulcer disease, atrophic gastritis and gastric cancer. Aliment Pharmacol Ther. 1997 Apr;11 Suppl 1:71-88. PMID:9146793
- ↑ Eaton KA, Brooks CL, Morgan DR, Krakowka S. Essential role of urease in pathogenesis of gastritis induced by Helicobacter pylori in gnotobiotic piglets. Infect Immun. 1991 Jul;59(7):2470-5. PMID:2050411
- ↑ Eaton KA, Krakowka S. Effect of gastric pH on urease-dependent colonization of gnotobiotic piglets by Helicobacter pylori. Infect Immun. 1994 Sep;62(9):3604-7. PMID:8063376
- ↑ Olson JW, Maier RJ. Molecular hydrogen as an energy source for Helicobacter pylori. Science. 2002 Nov 29;298(5599):1788-90. PMID:12459589 doi:10.1126/science.1077123
- ↑ Tomb JF, White O, Kerlavage AR, Clayton RA, Sutton GG, Fleischmann RD, Ketchum KA, Klenk HP, Gill S, Dougherty BA, Nelson K, Quackenbush J, Zhou L, Kirkness EF, Peterson S, Loftus B, Richardson D, Dodson R, Khalak HG, Glodek A, McKenney K, Fitzegerald LM, Lee N, Adams MD, Hickey EK, Berg DE, Gocayne JD, Utterback TR, Peterson JD, Kelley JM, Cotton MD, Weidman JM, Fujii C, Bowman C, Watthey L, Wallin E, Hayes WS, Borodovsky M, Karp PD, Smith HO, Fraser CM, Venter JC. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature. 1997 Aug 7;388(6642):539-47. PMID:9252185 doi:10.1038/41483
- ↑ Higgins KA, Carr CE, Maroney MJ. Specific metal recognition in nickel trafficking. Biochemistry. 2012 Oct 9;51(40):7816-32. doi: 10.1021/bi300981m. Epub 2012 Sep, 28. PMID:22970729 doi:10.1021/bi300981m
- ↑ Olson JW, Mehta NS, Maier RJ. Requirement of nickel metabolism proteins HypA and HypB for full activity of both hydrogenase and urease in Helicobacter pylori. Mol Microbiol. 2001 Jan;39(1):176-82. PMID:11123699
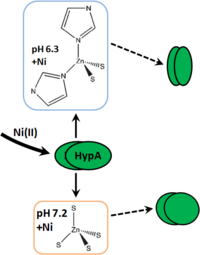
- ↑ 8.0 8.1 8.2 8.3 8.4 8.5 8.6 Herbst RW, Perovic I, Martin-Diaconescu V, O'Brien K, Chivers PT, Pochapsky SS, Pochapsky TC, Maroney MJ. Communication between the zinc and nickel sites in dimeric HypA: metal recognition and pH sensing. J Am Chem Soc. 2010 Aug 4;132(30):10338-51. doi: 10.1021/ja1005724. PMID:20662514 doi:10.1021/ja1005724
- ↑ Xia W, Li H, Sze KH, Sun H. Structure of a nickel chaperone, HypA, from Helicobacter pylori reveals two distinct metal binding sites. J Am Chem Soc. 2009 Jul 29;131(29):10031-40. PMID:19621959 doi:10.1021/ja900543y
- ↑ Watanabe S, Arai T, Matsumi R, Atomi H, Imanaka T, Miki K. Crystal structure of HypA, a nickel-binding metallochaperone for [NiFe] hydrogenase maturation. J Mol Biol. 2009 Dec 4;394(3):448-59. Epub 2009 Sep 19. PMID:19769985 doi:10.1016/j.jmb.2009.09.030