SP3.4-TCR-HLA-DQ8-α-1-gliadin complex
From Proteopedia
Introduction
Celiac disease is a disorder that damages the lining of the small intestine when dietary gluten such as wheat barley and rye is consumed. When a person with celiac disease ingests gluten an immune response engages and damages the microvilli of the small intestine. This damage prevents nutrient absorption and causes malnourishment. [1]Celiac disease is typically a predisposed genetic disorder in individuals that express a human leukocyte antigen (HLA) DQ2 and/or HLA-DQ8 molecules. Gluten peptidases bind to these HLA-DQ8 and HLA-DQ2 molecules and cause a T cell mediated response. Specifically, TRBV9*01 is the T-cell responsible for the recognition of the HLA-DQ8-∝-I-gliadin.[2]
Background Information
In patients with celiac disease the antigen is gluten, which consists of a mixture of gliadin and glutenin polypeptides. Proteins, which are usually degraded by gastric, pancreatic, and intestinal enzymes do not usually stimulate an immune response because they are too small too bind to HLA molecules. However, gluten is able to stimulate an immune response because gluten consists of a proline rich peptide and survives through the enzymes in the gastrointestinal tract. The peptide passes through the gastrointestinal lining into the subepithelial space.[3] In the subepithelial space an enzyme called TTG deamidates the gliadin peptide at β57 changing the amino acid from Asp to Ala.[4] This deamidation makes the peptide antigenic, which allows for the identification by HLA-DQ8 and/or HLA-DQ2, the autoantigens of celiac disease. Autoantigens are apart of the immune response defense mechanism of the body and are also known as human leukocyte antigens of the major histocompatibility complex. HLA molecules bind to specific antigens.
HLA-DQ8 and HLA-DQ2 are apart of the class II MHC molecules, whose activity is commonly associated with the development disease. Between 90-95% of patients with celiac disease express HLA-DQ2 and the remaining express HLA-DQ8. Individuals with the disease can express both human leukocyte antigens. [5]
In patients with celiac disease TRBV9*01 is the T-cell responsible for the recognition of the HLA-DQ8-α-I-gliadin. Multiple types of TRBV9*01 T-cells recognize the HLA-DQ8-α-I-gliadin complex. SP3.4 is an example of a TRBV9*01 T-cell that recognizes this complex.[6]
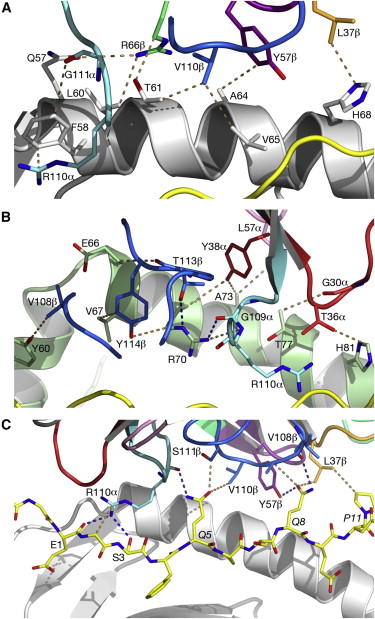
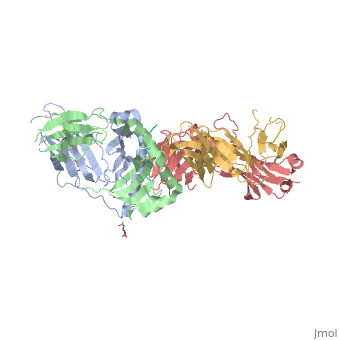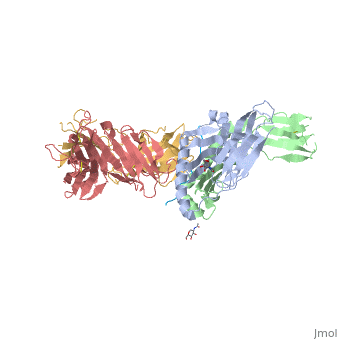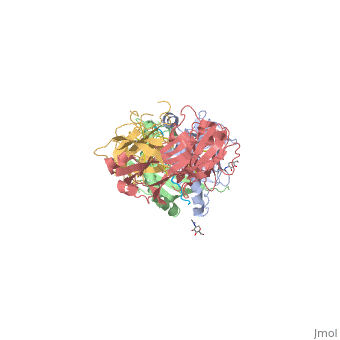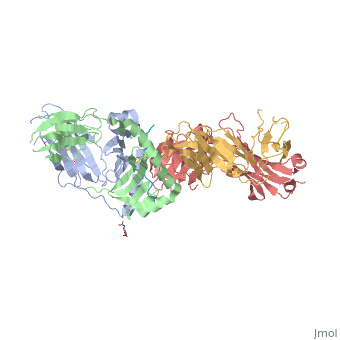
Structure of SP3.4 TCR-DQ8-Glia-αI ComplexThe SP3.4 T cell receptor interacts with the HLA-DQ8-glia-α1 at 70° across the long axis of the antigen binding cleft. At this antigen binding cleft, the Vα domain of the SP3.4 TCR sits directly above the β helix of HLA-DQ8 and the Vβ domain of the SP3.4 TCR sits directly above the α helix of HLA-DQ8. The buried surface area (BSA) of SP3.4 TCR-DQ8-glia-α1 interface is 880 Ų. This value BSA displays a standard docking mode to autoimmune TCR-pMHC-II. This indicates this disease is associatd with an autoimmune response and demonstrates how a non self antigen restricted to HLA-DQ8 is identified by SP3.4 TCR. The CDR loops of the SP3.4 TCR display limited plasticity in the process of docking to the HLA-DQ8-glia-α1 complex. On the SP3.4 TCR, the CDR1α loop moves 1.2Å away and CDR3α loop moves 2Å away to engage the HLA-DQ8-glia-α1 complex. The slight movement in these particular loops of CDRα may relate to cell signaling. However, the CDRβ loops do not change conformation to promote ligation. Val67 and Arg70 are the only side chains on the HLA-DQ8-glia-α1 complex that move to allow ligation with the SP3.4 TCR. Val67 and Arg70 orientate to avoid steric hindrance with Tyr114β and sequentially interact with the CDR3α and CDR3β loops. The ligation of SP3.4 TCR and HLA-DQ8-glia-α1 held together by ten hydrogen bonds and multiple van der Waals interactions. These van der Waals and hydrogen bond interaction between side chains allows for the centrally docking of SP3.4 TCR to HLA-DQ8 during ligation. The interaction between the α chain of the HLA-DQ8 and SP3.4 TCR is entirely due to van der Waals interactions. These van der Waals interactions are stabilized by the CDR3α, CDR1β, and CDR2β loops. The α chain of the HLA-DQ8 is composed of Phe58 to His68. Below the CDR3α loop Phe58 resides and packs against Arg110α due to its aromatic ring. Arg66β is adjacent to the CDR2β loop and interacts with Thr61 and Leu60. Tyr57β of the CDR2β loop packs against Ala64 and points towards His68. His68 interacts with Leu37, which resides in the CDR1β loop. The ligation of SP3.4 TCR and DQ8-glia-α1 held together by ten hydrogen bonds and multiple van der Waals interactions. These van der Waals and hydrogen bond interaction between side chains allows for the centrally docking of SP3.4 TCR to HLA-DQ8 during ligation. The interaction between the α chain of the HLA-DQ8 and SP3.4 TCR is entirely due to van der Waals interactions. These van der Waals interactions are stabilized by the CDR3α, CDR1β, and CDR2β loops. The α chain of the HLA-DQ8 is composed of Phe58 to His68. Below the CDR3α loop Phe58 resides and packs against Arg110α due to its aromatic ring. Arg66β is adjacent to the CDR2β loop and interacts with Thr61 and Leu60. Tyr57β of the CDR2β loop packs against Ala64 and points towards His68. His68 interacts with Leu37, which resides in the CDR1β loop. Arg70 resides across the antigen binding cleft and interacts with Gly109α and Thr113β of SP3.4 TCR via van der Waals contact and Hydrogen bonds. The conversion of Leu57α of the CDR2α loop with Thr36α and Tyr38α of the CDR1α loop form a hydrophobic focal point in the β1 helix. This hydrophobic focal point allows for Leu57α Thr36α and Tyr38α to interact with the HLA-DQ8 side chains His81, Ala73, and Thr77.[7]
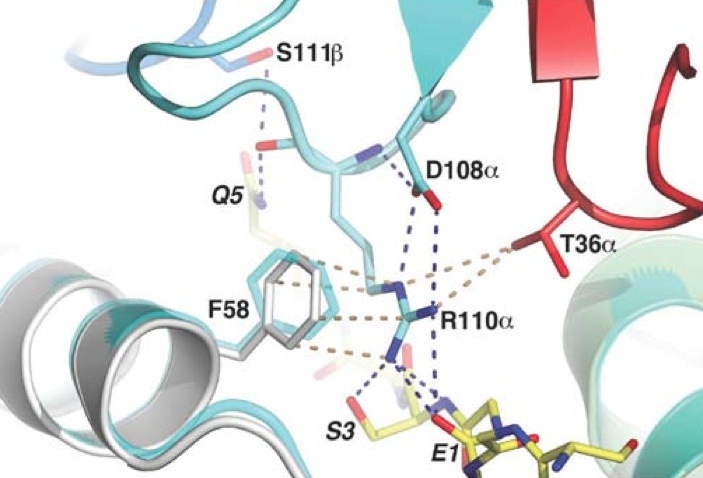
Interactions with α-I-Gliadin PeptideThe positions P1-Glu, P2-Gly, P3-Ser, P5-Gln, and P8-Gln of the α-I-gliadin peptide interact with the CDR3α, CDR1β, CDR3β and CDR2β loops. Since eight out of the ten Hydrogen bonds present at the SP3.4 TCR-HLA-DQ8-glia-αI interface are mediated by the α-I-gliadin peptide residues, the polar interactions with the peptide are the strongest energy contributors to the ligation of SP3.4 TCR to HLA-DQ8-glia-αI. Arg110α of the CDR3α loop has its side chain next to the α-I-gliadin peptide and toward its N terminus. By forming four Hydrogen bonds, the guanidinium functional group interacts with the peptide. The guanidinium functional group also interacts with Phe58α of HLA-DQ8. Before ligation the CDR3α loop is disordered. The stabilization of Arg 110α at ligation causesthe CDR3α loop to become ordered. Arg110α is also stablaized at ligation from an intra chain salt bridge to Asp108α and hydrogen bond with Asp108α to the backbone of Arg110α. The prominent residues on the gliadin peptide are P5-Gln and P8-Gln. The CDR3β loop is in between these two residues. P5-Gln interacts with the CDR3α and CDR3β loops of SP3.4 TCR. Within this interaction, the P5-Gln forms a hydrogen bond with Arg110α and interacts with Ser111β via van der Waals contacts. P8-Gln interacts with CDR1β, CDR3β and CDR2β loops of SP3.4 TCR. P8-Gln forms hydrogen bonds with the main chain of Val108β of the CDR3β loop and with Tyr57β of the CDR2β loop. P8-Gln also form van der Waals contacts with Ala109β of the CDR3β loop and Leu37β of the CDR1β loop.[8]
SymptomsWhen a person with celiac disease ingests dietary gluten some of the symptoms include abdominal pain, bloating, indigestion, constipation, decreased appetite, nausea, vomiting, unexplained weight loss, depression, anxiety, fatigue, hair loss, itchy skin, missed menstrual periods, muscle cramps and joint pain, nosebleeds, and seizures.[9] TreatmentCurrent treatment for Celiac disease is to stay away from all gluten products. Increasded mortality due to cancer, infertility in women, decreased growth due to decrease in bone mineral are all long term symptoms of celiac disease if it is left untreated.[10]
Biological SignificanceIndividuals with the disease can express both human leukocyte antigens. HLA-DQ8 and HLA-DQ2 are necessary for disease development but not sufficient for development. For this reason, a person expressing these HLA can develop celiac disease at any point in their life or not at all. The development of celiac disease is still not clear. However by studying the mechanisms such as the TRBV9*01-TCR-DQ8-glia-α1 can help decipher the causes of celiac disease. This understanding can lead to a cure or prevention of the disease. For further study it would be beneficial to look at the interactions within the subepithelial space since this is the place where the gluten polypetides are deamidated and targeted as an antigen.[11]
References
|
| |||||||||||