From Proteopedia
proteopedia linkproteopedia link T7 RNA Polymerase Initiation Complex
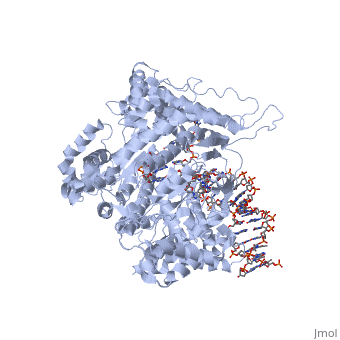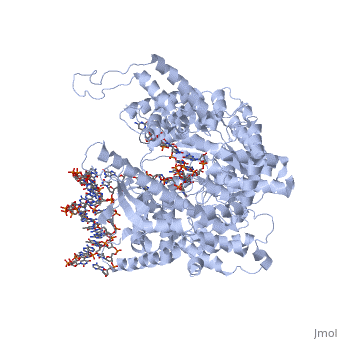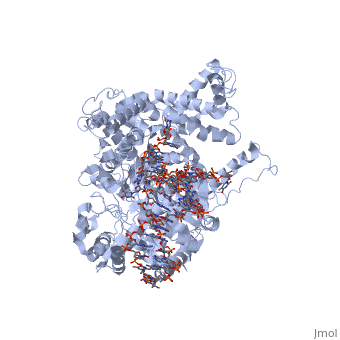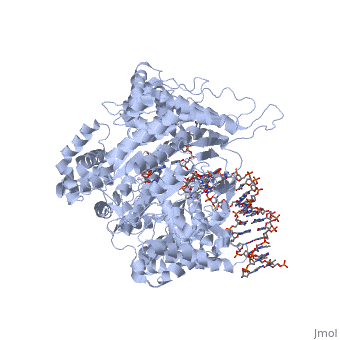
| Promoter binding. T7 RNA polymerase , by recognizing the of the promoter (-17 to -5), and then melting a bubble (-4 to about +3), within the larger duplex. The duplex promoter domain binds primarily to the N-terminal domain of the enzyme, with the exception of the (C-terminal domain) "." It is the combination of the N-terminal domain, with the positioned specificity loop, that forms the specific binding surface.
Intercalating loop stabilizes the melted complex. Strong interactions with the duplex region of the promoter places the "" into the DNA between residues -4 and -5. The intercalating loop, also called the Valine Loop, has hydrophobic residues Val, Ile, etc that stack on and stabilize the exposed face of the base pair at position -5, stabilizing the locally melted structure.
Positioning of the +1 and +2 bases in the active site. Melting of a bubble within the DNA allows the (single stranded) template strand to enter the active site, and allows template strand bases +1 and +2 to orient in the active site, poised for initiation. Note that GTP at position +2 is in the normal substrate position and is that will be used in catalysis. GTP at position +1, by contrast, sits where the 3' base of the elongating RNA normally sits. In an elongation complex, that base is held in place by the upstream duplex. During initiation, the , but that Mg(II) is not coordinated by the protein, so there is little binding stabilization. For this reason, Km for the +1 base is much higher (binding is weaker) than for all other (elongating) bases.
Catalysis. The enzyme then , as directed by the template (typically two GTP's, encoded by CC in the template strand). The to initiate a phosphoryl transfer reaction. Release of pyrophosphate (PPi) leaves the product dinucleotide (pppGpG) in the active site. Note that one of the the trigonal bipyramidal reaction intermediate (not shown) in this SN2 phosphoryl transfer reaction.
Movement. At this point, the complex is in the and to add the next base, the enzyme must translocate forward along the DNA (or equivalently, the RNA/DNA slides backwards), forming the post-translocated state. In the latter state (only) the active site now accommodates binding of the next NTP to the (+3) template base, to then .
This cycle of NTP binding, catalysis (bond formation via phosporyl transfer), and forward translocation repeats over and over, throughout extension of the RNA.
The enzyme active site presumably stabilizes these short hybrids, but evidence also suggests that the intercalating loop, upstream and the active site, downstream, stabilize the bubble and keep it from collapsing and competitively displacing the short, nascent RNA.
The initial bubble grows. From 2mer, the system progresses ->3mer->4mer->5mer->6mer ... During this time, the newly formed RNA-DNA duplex (hybrid) grows, from 2 bases, to , to 4 bases, to 5 bases, to 6 bases, to, to , etc., and during the time, the duplex is short and otherwise unstable.
The growing hybrid induces protein domain movement. Also note that the initial active site accommodates only about a 3 basepair RNA-DNA duplex, as the N-terminal domain lies in the path of that hybrid (remember that forward translocation of the polymerase is really reverse translocation of the RNA-DNA hybrid). Beyond about 3 bases, the , inducing into to both translate backwards and rotate. This can be seen in structures of the complex with 7 and 8 nucleotides of RNA synthesized. In the morph shown [1], models of how the structure might look with 4, 5, and 6 nucleotides of RNA synthesized were interpolated from the existing structures with 3 and 7 nucleotides of RNA. In this , the growing hybrid is shown explicitly (with some modeling problem for 6 nucleotides if you look closely).
Transition to Elongation. Toward the end of the above rotation (at about a 9mer RNA), stress builds up in the promoter binding domain, leading to a weakening of some of the interactions with the upstream duplex promoter. This triggers promoter release, which now allows the N-terminal domain to rotate 220° in the other direction, to form the .
Function
Disease
Relevance
Structural highlights
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.
|
Files for 3D printer
T7 RNA polymerase model by Marius Mihasan
References
1,"1aro","Free enz plus lysozyme","T7RP with the bound inhibitor T7 lysozyme, no DNA - Jeruzalmi, D. & Steitz, T. A. (1998) EMBO J 17, 4101-4113"
2,"1cez","Enz with DNA bound (ED complex)","Early structure of T7RP with promo bound - Cheetham, G. M., Jeruzalmi, D. & Steitz, T. A. (1999) Nature 399, 80-83"
3,"2pi5","Enz with DNA bound (ED complex)","T7RP with promoter and first two NTPs bound - Kennedy, W.P.,††Momand, J.R.,††Yin, Y.W. (2007) Mechanism for de novo RNA synthesis and initiating nucleotide specificity by t7 RNA polymerase. J.Mol.Biol. 370: 256-268"
4,"2pi4","ED complex with GTP + GTP","T7RP with promoter and first two NTPs bound - Kennedy, W.P.,††Momand, J.R.,††Yin, Y.W. (2007) Mechanism for de novo RNA synthesis and initiating nucleotide specificity by t7 RNA polymerase. J.Mol.Biol. 370: 256-268"
5,"1qln","ED with 3mer RNA","T7RP with promoter DNA and GTP, allowing formation of a 3 base transcript - Cheetham, G. M. & Steitz, T. A. (1999) Science 286, 2305-2309",true
7,"3e2e","Initial complex at +7","The structure of a transcribing T7 RNA polymerase in transition from initiation to elongation - Durniak, K.J., Bailey, S., Steitz, T.A. (2008) Science 322, 553-7"
6,"3e3j","Initial complex at +8","The structure of a transcribing T7 RNA polymerase in transition from initiation to elongation - Durniak, K.J., Bailey, S., Steitz, T.A. (2008) Science 322, 553-7"
8,"1msw","Elongation complex (Steitz)","Elongation complex model formed with mismatch bubble DNA - Yin, Y. W. & Steitz, T. A. (2002). Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase. Science 298, 1387-1395."
9,"1h38","Elongation w scaffold","Elongation complex model formed by multi-piece scaffold - Tahirov, T. H., Temiakov, D., Anikin, M., Patlan, V., McAllister, W. T., Vassylyev, D. G. & Yokoyama, S. (2002) Nature 420, 43-50"
10,"1s0v","Elongation w ab-me-ATP","Scaffold elongation complex with non-hydrolyzable substrate NTP - Temiakov, D., Patlan, V., Anikin, M., McAllister, W. T., Yokoyama, S. & Vassylyev, D. G. (2004) Cell 116, 381-391"
11,"1s76","Elongation w ab-me-ATP","Mismatched bubble elongation complex with non-hydrolyzable substrate NTP - Yin, Y. W. & Steitz, T. A. (2004) Cell 116, 393-404"
12,"1s77","Elongation w PPi","Yin, Y. W. & Steitz, T. A. (2004) Cell 116, 393-404"
13,"4rnp","Low res free enz"
- ↑ The Storymorph Jmol scripts were used to create the interpolation shown in the morph. Coordinates available on Proteopedia