User:Brittany Allen/Sandbox 1
From Proteopedia
|
Contents |
General Background Information
TIP30, also known as both CC3 [1] and HTATIP2 [2], is a tat-interacting protein that functions as a tumor suppressor involved in cell growth, apoptosis, metastasis, DNA repair, metabolism and angiogenesis of tumor cells [2,3,4]. TIP30 acts as a transcription cofactor that can regulate gene expression [5] and has both pro-apoptotic and anti-metastatic properties [6]. When the promoter of TIP30 is methylated, TIP30 becomes downregulated and associated with tumor prognosis[2]. It is thought that the tumor suppressor effect is the result of the inhibition of nuclear transport through binding with importin βs or by regulating transcription through interaction as a complex with a co-activator independent of AF-2 function and the c-MYC gene[7]. Several studies have found that TIP30 may be linked to esophageal carcinoma, laryngeal carcinoma, glioma, pancreatic ductal adenocarcinoma, breast cancer, gastric cancer, gallbladder adenocarcinoma, lung cancer, and hepatocellular carcinoma[1].
Structure
TIP30 has a molecular mass of 30 kd[5] and is composed of 242 amino acids[1]. TIP30 is composed of alpha helices, beta sheets, and loops as seen in Image 1. A study looking into the structure of TIP30 has found that it is dominated by a dinucleotide (Rossman) fold located in the N terminal two-thirds of TIP30[6].
Through sequence analysis studies, it is thought that TIP30 may be a member of the SDR (substrate determining residue) family which contain a characteristic motif at their catalytic start sites [1]. The carboxyl terminus of TIP30 binds to the SDR substrate, while the amino terminus of TIP30 is the nucleotide cofactor-binding domain which has a characteristic Gly-X-X-Gly- X-X-Gly motif (where X can be any amino acid) [1]. Since SDR families have binding specificity for NADPH [8] and TIP30 contains a dehydrogenase reductase fold that contains binding specificity for NADPH [1], the binding of NADPH may be important for the biological activity of TIP30 including interactions with importins as well as the c-MYC system[8].
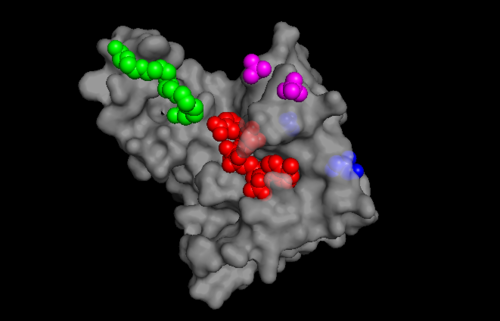
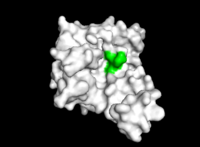
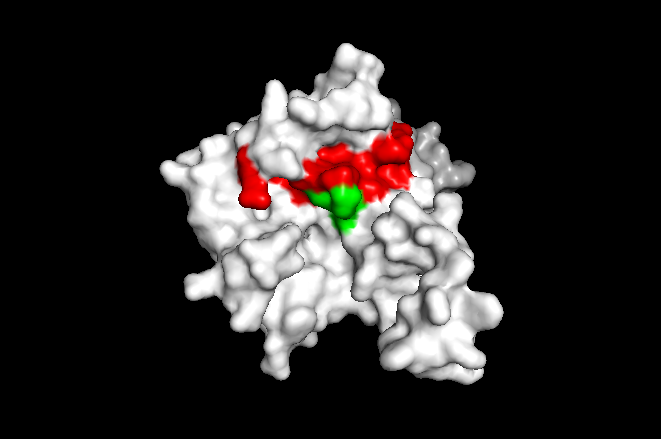
According to Uniprot, TIP30 contains a nucleotide binding region between residues 19-52, as shown in image 3 and a binding site at residue 131. When comparing images 2 and 3, it can be observed that the ligands bind in the active regions and interact with the residues. Uniprot also noted that TIP30 can contain a mutagenic site at positions 28-31 (image 4), if this site is present there is a loss of proapoptotic and metastasis-inhibiting effects.
Function
TIP30 is found in both the cell nucleus and cytoplasm [10] and functions in a variety of cellular processes including apoptosis, proliferation, metastasis, angiogenesis, DNA damage repair, and metabolic adaptation [1]. In normal cell growth, TIP30 regulates DNA replication and repair. Stress activates TIP30 to regulate cell proliferation through cell cycle arrest, senescence, and apoptosis to prevent tumor formation. In this manner, TIP30 can act as a tumor-suppressor gene of Type I (caretaker) or a Type 2 (gatekeeper) to prevent mutagenesis [1]. Type I tumor-suppressor genes work by repairing damaged DNA while Type 2 halt the cell cycle to allow time to fix the DNA [1].
TIP30 Tumor Suppressive Functions
TIP30 regulates apoptosis
TIP30 has pro-apoptotic activity[1]. When normal cellular levels of TIP30 are increased, the excess TIP30 inhibits nuclear transport both in vitro and in vivo[10]. TIP30 will bind to the karyopherins[6] of the importin beta family in a RanGTP-insensitive manner and associate with nucleoporins[10]. This action will inhibit the nuclear import of proteins possessing nuclear localization signals[10]. When TIP30 forms this complex with RanGTP with importing, it will induce apoptosis [10].
TIP30 regulates proliferation
In a study done with HCC cells, when TIP30 is expressed with increased levels of p53, and decreased levels of Bcl-2, and combined with cytotoxic drug 5-fluorouracil treatment, tumor growth was suppressed, implicating TIP30 to play a role in suppressing tumor proliferation[1].
TIP30 regulates metastasis
TIP30 can inhibit metastasis through two mechanisms: one is through an apoptosis-inducing effect, and second through an angiogenesis-inhibiting effect[1]. TIP30 is able to inhibit the expression of tumor metastasis-associated genes including osteopenia and calcium adhesion proteins[3].Studies suggest that TIP30 may actually use signaling through EGFR to inhibit metastasis in certain cancers [6].
TIP30 regulates angiogenesis
Angiogenesis is the process of cells sprouting new capillaries from existent vessels in order to help tumor growth[11]. In a tumor, the cells will secrete fibroblast growth factors (aFGF and bFGF), vascular endothelial cell growth factors (VEGF), and antiangiogenic factors including thrombospondin, angiostatin, and endostatin that influence whether or not the tumor will be able to grow and replicate[11]. However, in vitro experiments have demonstrated that TIP30 can induce changes in RNA levels of angiogenic modulators that reduce angiogenesis[1,11]. In an in vitro experiment using both macro and microvascular origin tumor cells, angiogenic stimulator angiopoietin I (a pro-angiogenic protein) was reduced while angiogenic inhibitors had increased expression, indicating that TIP30 uses angiogenic modulators to inhibit angiogenesis[1].
TIP30 controls gene expression
TIP30 is a gene transcription factor that uses tat proteins and RNA polymerase II[1]. TIP30 can mediate tumor suppressive regulation by directly affecting the transcription of genes in the nucleus or by indirectly changing the signal transduction pathways in the cytoplasm or inhibiting nuclear import of proteins[1]. TIP30 also plays a role in the regulation of EGFR, a protein that plays a role in the pathogenesis of human cancers[1].
TIP30 regulates tumor cell metabolism
In an experiment with silenced TIP30 in HeLa cells, the researchers were able to demonstrate that in the silence of TIP30, tumor cells can thrive in a low glucose setting[1]. When TIP30 is absent, mitochondrial oxidative phosphorylation of glucose is low[1]. The silencing of TIP30 in tumor cells is dependent on glycolysis to give the cells flexibility to use both OXPHOS and glycolysis for metabolism[1].
TIP30 regulates DNA damage responses
Following DNA damage from ultraviolet light and oxidant exposure, TIP30 can regulate the metabolic adaptation of the cells with glucose limitations[2]. The TIP30 can then bind with p53 and regulate the BAX gene which sensitize the cells to apoptosis[2].
Role in Disease
TIP30 is naturally expressed in human tissues including the heart, brain, lung, kidney, and pancreas. However, in cancers-- including breast cancer, lung cancer, liver cancer, gastric cancer, and colorectal cancer, TIP30 expression is decreased. As shown in the section above, many of the functions of TIP30 play a critical role in the prevention of disease through the control of apoptosis, growth, metastasis, angiogenesis, DNA repair, and tumor cell metabolism. When TIP30 is properly expressed, patients will have a good prognosis[2],but when the TIP30 promoter becomes methylated[2,4], or TIP30 becomes lost or decreased in expression, this can contribute to irregular cell growth and development and cause tumors[1]. For example, high levels of TIP30 can be beneficial and induce apoptosis, meanwhile the down regulation of TIP30 in tumor cells leads to further growth and development of tumors[3].
Research has shown that the silence of TIP30 improves the survival of tumor cells as a response to glucose limitation[12]. One experiment focused on the silence of TIP30 in HeLa cells[12]. With TIP30 silenced, the cell was able to maintain high levels of mitochondrial respiration, allowing them to survive on little glucose[12]. The cells also experienced a higher expression of mitochondrial proteins in respiration complexes, as well as increased levels of c-MYC and the M2 isoform of pyruvate kinase in the low glucose setting[12]. Thus, in the absence of TIP30, the cells were still able to operate and function in a low glucose environment which allowed the cells to continue to multiply and grow the tumor. Additional studies have demonstrated that iIf TIP30 is mutated in the nucleotide binding motif (between residues 25-31[6], or resides 28-21 [uniprot]), the cell will lack apoptic properties and have weakened importing interactions[6]. The mutagenesis site can be seen in image 4 where the mutated residues are colored green.
In another recent study, the expression of TIP30 was researched in patients with cancer. Over 14 studies, that included 1705 patients, an association was observed that linked TIP30 expression to the good overall survival of the cancer patients[5]. Due to this study, along with previous research, promoting the expression of TIP30 is a solid potential for drug targets because if TIP30 is expressed, and not down regulated, you could prevent one source of forming tumors[5].
Evolutionary Conservation
TIP30 is an evolutionary conserved gene located on human chromosome 11 [1]. When analyzing the amino acid sequence of TIP30, 98% of the protein was identical to CC3 and after further analysis, TIP30 and CC3 were identified to be the same protein[1].
A BLAST analysis using the FASFA sequence of TIP30 from Uniprot [13], revealed that TIP30 has sequence similarity to SDR superfamilies. Further analysis revealed that TIP30 also contains several NAD(P) binding domains [13]. The domains that were shown to be found in the TIP340 sequence can be seen in image 6. One study indicated that TIP30 has the closest match to UDP-galactose epimerase with ~17.5% sequence identity[6]. Another studied revealed that TIP30 was ~22% similar to the cAMP-dependent protein kinase catalytic subunit of Caenorhabditis elegans.[6]
References
1.Yu X, Li Z, Wu WKK. TIP30: A Novel Tumor-Suppressor Gene. Oncology Research Featuring Preclinical and Clinical Cancer Therapeutics [Internet]. 2015;22(5):339–48. Available from: http://www.ingentaconnect.com/content/cog/or/2015/00000022/F0020005/art00013#
2.Xu T, Jin Z, Yuan Y, Zheng H, Li C, Hou W, et al. Tat-Interacting Protein 30 (TIP30) Expression Serves as a New Biomarker for Tumor Prognosis: A Systematic Review and Meta-Analysis. Plos One [Internet]. 2016;11(12). Available from: http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0168408
3.Hu Y, Chen F, Liu F, Liu X, Huang N, Cai X, et al. Overexpression of TIP30 inhibits the growth and invasion of glioma cells. Molecular Medicine Reports [Internet]. 2015;13(1):605–12. Available from: https://www.spandidos-publications.com/mmr/13/1/605?text=fulltext
4.Fong S, King F, Shtivelman E. CC3/TIP30 affects DNA damage repair. BMC Cell Biology [Internet]. 2010;11(1):23. Available from: https://bmccellbiol.biomedcentral.com/articles/10.1186/1471-2121-11-23
5.Zhang X, Lv L, Ouyang X, Zhang S, Fang J, Cai L, et al. Association of TIP30 expression and prognosis of hepatocellular carcinoma in patients with HBV infection. Cancer Medicine [Internet]. 2016;5(9):2180–9. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5055146/
6.Omari KE, Bird LE, Nichols CE, Ren J, Stammers DK. Crystal Structure of CC3 (TIP30). Journal of Biological Chemistry [Internet]. 2005;280(18):18229–36. Available from: http://www.jbc.org/content/280/18/18229.long
7.Jörnvall H. Medium- and short-chain dehydrogenase/reductase gene and protein families. Cellular and Molecular Life Sciences [Internet]. 2008;65(24):3873–8. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2792337/
8.K EO, LE B, CE N, J R, DK S. K [Internet]. PDB 2bka structure summary ‹ Protein Data Bank in Europe (PDBe) ‹ EMBL-EBI. 1970 [cited 2017Dec4]. Available from: https://www.ebi.ac.uk/pdbe/entry/pdb/2BKA
9.European Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. Oxidoreductase HTATIP2 [Internet]. HTATIP2 - Oxidoreductase HTATIP2 - Homo sapiens (Human) - HTATIP2 gene & protein. 2017 [cited 2017Dec4]. Available from: http://www.uniprot.org/uniprot/Q9BUP3
10.King FW, Shtivelman E. Inhibition of Nuclear Import by the Proapoptotic Protein CC3. Molecular and Cellular Biology [Internet]. 2004;24(16):7091–101. Available from: https://www.ncbi.nlm.nih.gov/pubmed/15282309
11.Nicamhlaoibh R, Shtivelman E. Metastasis suppressor CC3 inhibits angiogenic properties of tumor cells in vitro. Oncogene [Internet]. 2001;20(2):270–5. Available from: https://www.nature.com/articles/1204075
12.Chen V, Shtivelman E. CC3/TIP30 regulates metabolic adaptation of tumor cells to glucose limitation . Cell Cycle [Internet]. 2010;9(24):4941–53. Available from: https://www.ncbi.nlm.nih.gov/pubmed/21150275
13.European Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. [Internet]. HTATIP2 - Oxidoreductase HTATIP2 - Homo sapiens (Human) - HTATIP2 gene & protein. 2017 [cited 2017Dec5]. Available from: http://www.uniprot.org/uniprot/Q9BUP3
14.NCBI Conserved Domain Search [Internet]. National Center for Biotechnology Information. U.S. National Library of Medicine; [cited 2017Dec5]. Available from: https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi
![Image 1. The secondary structures found in TIP30 as seen on PDB[8]. Red/pink resemble alpha helices, yellow arrows represent beta sheets, and green represent loops.](/wiki/images/thumb/a/a3/PDBstructure.png/700px-PDBstructure.png)

![Image 6. Using the FASFA sequence of TIP30 in a NCBI search for conserved domains revealed the presence of YbjT, NAD binding, And, and SDR domains[14].](/wiki/images/thumb/4/43/BLAST2_seq.png/600px-BLAST2_seq.png)
