This is a default text for your page Felipe de Melo Santana/Sandbox 1. Click above on edit this page to modify. Be careful with the < and > signs.
You may include any references to papers as in: the use of JSmol in Proteopedia [1] or to the article describing Jmol [2] to the rescue.
General information
OxyR is a protein that functions as a transcription factor. The OxyR we are analyzing is that of Escherichia Coli. In this perspective, the protein works by regulating the transcription of specific genes, being sensitive to H2O2.Thus, OxyR acts mainly as a transcriptional activator under oxidizing conditions through direct interaction with the RNA polymerase a subunit.This transcription factor belongs to the Lysr family, with 34 kDa, which forms homotetramers. Under this focus, the members of this family present a domain linked to DNA in their N-terminal region, called helix-loop-helix, a central domain responsible for recognizing H2O2 that is sensitive to the regulation signal and, finally, a domain in C-terminal region responsible for activation and mutimerization. it can be concluded that the OxyR gene encodes a protein that has a dual function, acting as a stress sensor and also as a transcription activator of the OxyR operon.
Structure
This protein its a transcription factor, at that time its shows two different structures when its reduced or oxidized. The reduced form monomer of the OxyR regulatory domain consists of two α/β domains that are linked by two interdomain strands. Residues in the middle of a folded domain form the majority of the N-terminal a/b domain. The C-terminal a/b Results and Discussion domain II is made of residues .
The remaining C-terminal 30 Structure Determination residues come back to domain I to form an edge of the domain . The two domains exhibit a similar folding pattern consisting of a central b sheet flanked by helices and loops.
The oxidized form is characterized by the ediction of the domain II, resulting in a significant rearrangement of secondary structures.
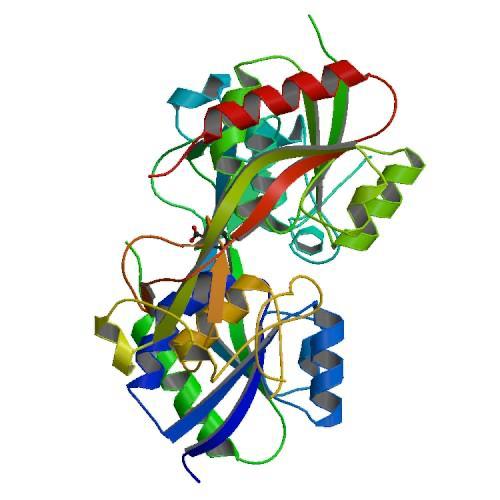
Figure 1. CRYSTAL STRUCTURE OF THE REDUCED FORM OF OXYR (1I69)
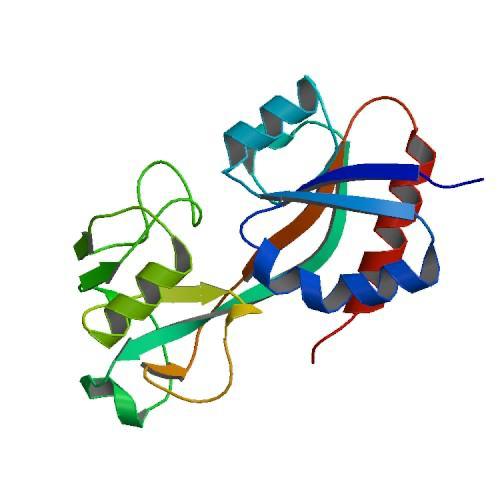
Figure 2. CRYSTAL STRUCTURE OF THE OXIDIZED FORM OF OXYR (1I64)
OxyR activation
The OxyR transcription factor is activated by the formation of an intramolecular disulfide bond (Zheng et al. 1998).
The OxyR activation, by H2O2, begins with the oxidation of the residue into a sulfenic acid intermediate, which causes destabilization of the Cys199 side chain in the hydrophobic binding pocket, due to its increased size and charge. The destabilization leads to expulsion of the side chain of Cys199 out of the interdomain pocket, resulting in a flexible loop around Cys199 (Lee et al. 2004).
This change in the regulatory domain of OxyR allows the formation of an intramolecular disulfide bond between , responsible for keeping OxyR in the active form. Thus, OxyR is deactivated by the reduction of the disulfide bond, that leads OxyR back to its reduced form (Zheng et al. 1998).
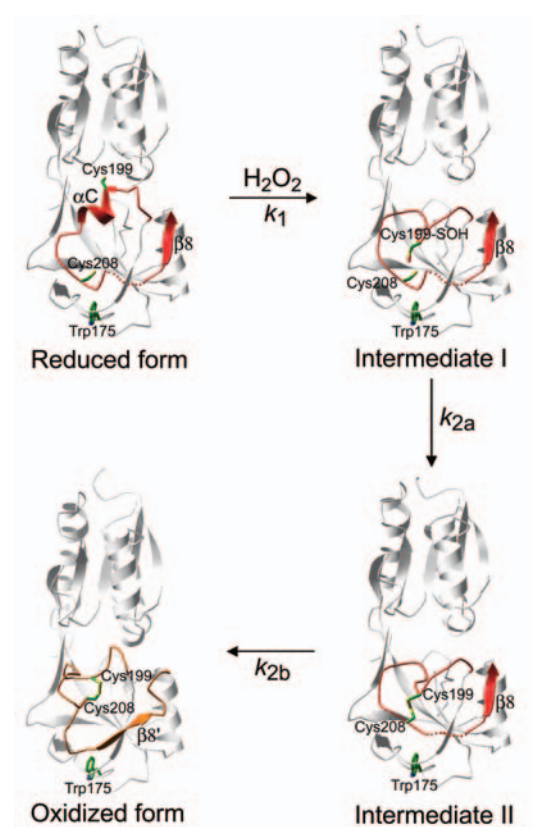
Figure 3. Intermediates in the structural transitions of OxyR. Lee et al. 2004
Protein activity in different forms
The OxyR transcription factor regulate the expression of genes involved in redox-homeostasis (Stroz et al. 1990).
OxyR protein is able to DNA-binding in its two different forms, oxidized and reduced. In your oxidized form, OxyR binds to four adjacent major grooves in DNA, while in the reduced form, OxyR binds to two pairs of adjacent major grooves separated by one helical turn (Toledano et al 1994). These two modes of binding allow OxyR to regulate different promoters, under different cell conditions.
Both forms of OxyR act as repressors, regulating its own expression by oxyR transcription repression, while only oxidized OxyR is able to activate gene expression, inducing the expression of oxyS (a small regulatory RNA), hydroperoxidase I (katG), alkyl hydroperoxide reductase (ahpCF), glutathione reductase (gorA), glutaredoxin 1 (grxA) and other genes involved in redox-homeostasis, that protect the cell against oxidative stress (Zheng et al. 1998).
Figure 4. Tetrameric Structure and a Plausible Model of DNA Interactions. Choi et al. 2001
The tetramers observed in crystals of the reduced (A) and oxidized (B) forms are shown with the DNA positioned on the tetramers with plausible orientations. The tetramers are shown as ribbons (red, green, blue, and yellow for each monomer) and the model of DNA is represented as helical coils (cyan).
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.




