User:Luke Edward Severinac/Sandbox 1
From Proteopedia
Caspase-6 in Homo sapiens
Caspase-6 is an endopeptidase involved in apoptosis. In terms of its catalytic function, it is a part of the cysteine-aspartate family. Before Caspase-6 becomes functional, the enzyme exists as a , also known as a zymogen. This zymogen exists as a , whose are then cleaved at to assume its active conformation. Zymogen activation through cleavage is largely conserved across the caspase family. However, Caspase-6 is unique in that it becomes active through self-cleavage in addition to cleavage by a separate enzymes[1]. Each monomeric unit of zymogen contains a consisting of two helices, a consisting of three helices, a , and a . After cleavage at all sites, the processed post-zymogen monomers remain closely associated together through intermolecular forces as a dimer.
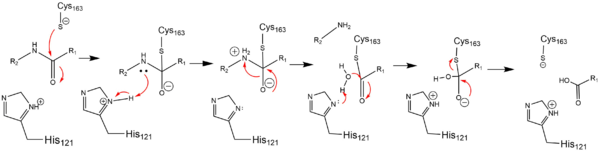
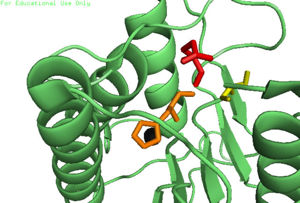
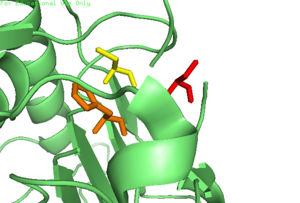
ZymogenIn addition to a self-cleavage mechanism, Caspase-6 can be activated through cleavage by Caspase-3, as well as other enzymes. This activation by cleavage is highly conserved across the caspase family, but activation through self-cleavage is uniquely recognized as the primary mechanism for Caspase-6 activation. In this self-cleavage mechanism, cleavage must occur at in order to remove the located at the N-terminus and the located within the protein. These cleavages are both sequence specific and ordered, starting with cleavage of the pro-domain at . Removal of the intersubunit linker then occurs through cleavage at two sites, [2]. It has been proposed that this sequence of cleavage is due to the pro-domain being more readily available to enter the active site, whose presence inhibits Caspase-6's ability to cleave the intersubunit loop and self-activate; The prodomain acts as a “suicide protector”, preventing the TEVD193 cleavage site from the active site[3]. After both cleavages occur, remains in solution as a dimer. Active StateIn order to function as an endopeptidase, each of active Caspase-6 utilizes a composed of , , and to cleave polypeptide ligands that can include neuronal proteins and tubulins[4]. In the theorized mechanism, atoms are shown in their resting ionic states; His-121 acts as an acid catalyst, Glu-123 acts as a base catalyst to deprotonate Cys-163, which then acts as covalent catalyst. Zinc InhibitionCaspase-6 can also assume an inactive state, which exists as a in its biological unit. For each , Caspase-6 function is primarily inhibited by the binding of a ion, which binds to an instead of the . This allosteric site is located on the opposite side of the protein relative to the active site. The zinc ion is bound to , Lys-36, Glu-244, and His-287. Once the ion is bound to the protein, it is then stabilized by a found in the cytoplasm. The binding of zinc at the exosite is suggested to cause a conformational change in the protein from an to an that misaligns catalytic residues and inhibits activity of the enzyme. It has been proposed that helices of the active dimer must rotate or move in some other way to provide these ideal interactions with zinc. This subtle shift is most likely the cause for allosteric inhibition[1]. As the helices move to bind zinc, the amino acids of the active site become misaligned. The altered positions of the amino acids no longer provide ideal interactions for incoming substrates. After zinc binds, substrates may still enter the active site, but no catalytic activity will occur. The first image shows the catalytic triad of Caspase-6 with zinc bound, and the second image shows the catalytic triad of caspase-6 without zinc bound. The catalytic cysteine and glutamate residues flip positions and become misaligned resulting in a loss of enzymatic function. PhosphorylationThe function of Caspase-6 can be inhibited by phosphorylation of Ser-257. The exact mechanism of this reaction remains unidentified at the time of publication, but proceeds when ARK5 kinase is present. This modification can occur before and after zymogen activation. The phosphoryl group inhibits Caspase-6 through steric interference. When Ser-257 is phosphorylated, the amino acid residue interacts with , causing a shift in the helices of Caspase-6[3]. This is shown in the , whose mutation mimics phosphorylation[5]. The shift misaligns and disrupts residues found in the active site. This conformational difference prevents the intersubunit linker from entering during zymogen activation and the self-cleaved active dimer cannot be formed. Additionally, no new substrate is able to enter the active site. Medical RelevanceCaspase-6 involvement in Alzheimer's DiseaseCaspase-6 is known to be involved in many neurodegenerative diseases, one of which is Alzheimer's disease (AD). Caspase-6 activity is associated with the formation of lesions within the Alzheimer's Disease.Lesions can be found in early stages of AD. A proapoptotic protein, p53, is present at increased levels within AD brains, which seems to directly increase the transcription of Caspase-6, which indirectly influences apoptosis of neurons. Future treatments of AD include selective inhibition of active Caspase-6 proteins; staining has found active Caspase-6 within the hippocampus and cortex of the brain within a varying severity of AD cases. This suggests that Caspase-6 plays a predominate role in the pathophysiology of AD. There has been research conducted that shows activation of Caspase-6 in AD could cause disruption of the cytoskeleton network of neurons and lead to neuronal apoptosis[3]. References
| ||||||||||||