Introduction
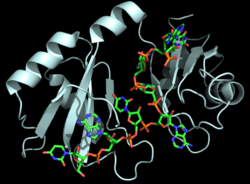
Figure 1: Cartoon representation of the Hrp1-PEE complex. The RNA is shown as a stick model and is colored by element.
Hrp1 is a polyadenylation factor found in Saccharomyces cervisiae (yeast) [1]. Hrp1 specifically recognizes and binds to an RNA sequence in the 3'UTR of the messenger RNA (mRNA) upstream from the cleavage site called the polyadenylation enhancement element (PEE) (Figure 1) [1]. Upon binding to the RNA, Hrp1 helps recruit additional proteins necessary for the cleavage and polyadenylation of the RNA molecule [1]. Although Hrp1 shares several common features with other RNA-binding proteins, the unique structural features of the Hrp1-PEE complex reveals the mechanism by which Hrp1 is able to recognize and bind to its specific RNA sequence at the atomic level [1]. Hrp1 was discovered when Cleavage Factor I (CF I) was purified and separated into its two components, CF IA and CF IB. CF IB is a single 73 kDa polypeptide. The polypeptide was digested and two tryptic peptides were obtained for sequencing. The sequences were aligned via a database, and Hrp1 was determined to be a perfect match. Hrp1 of CF IB interacts with Rna14 and Rna15 of CF IA[2] to form a protein complex that aids in cleavage, polyadenylation, and transport of the mRNA from the nucleus[3].
Structure
General Features
Hrp1 is a single strand RNA-binding protein composed of two RNP-type RNA-binding domains (RBDs) arranged in tandem with a typical ßαßßαß architecture [1]. The two RBDs have similar topolgies, both containing a central antiparallel four-stranded with two α-helices running across one face [1]. The two RBDs associate to form a deep and positively charged , which constitutes the binding site for the RNA molecule [1].
Hrp1-RNA Interactions
The interface between Hrp1 and its target RNA sequence is dominated by interactions between key aromatic residues and RNA nucleobases [1]. Only six RNA bases, an repeat, act as the PEE and form specific contacts with Hrp1 [1]. The kinked conformation around Ade4 is uncommon for RNA alone, and may be adopted by the RNA for specific interactions with Hrp1. Ade4 is part of a crucial interaction with Trp168 which will be discussed later, and could explain the adoption of the kinked conformation. Hydrophilic residues of Hrp1 provide base specificity through hydrogen bonding [1]. Most of the key residues that interact with the RNA can be found in the ß-sheet region of Hrp1; however, loops and the interdomain linker are also essential for Hrp1-RNA recognition [1]. Perhaps the most important Hrp1-RNA interaction is the (a conserved residue) [1]. In this case, Trp168 stacks on Ade4 and forms crucial base-specific hydrogen bonds [1]. It is also worth noting that a second Hrp1 residue is critical to holding Ade4 in place, , which stabilizes Ade4 likely via a cation–π interaction. A third contributor, , also stacks with Ura7 to aid in RNA recognition and binding [1].
RBD-RBD Interactions and the Linker Region
As mentioned above, Hrp1 is composed of two RBDs. The RBDs are connected by a (a short two-turn α-helix) which also contains an crucial residue for RNA binding. Ile234 holds Ade6 stacked in place with Phe162 . Experimental evidence from the NMR data [1] suggests that the two RBDs at independently until binding the PEE. Binding the PEE causes the linker region to adopt a short helical structure to rigidly hold the . Aside from the linker helix, the only interaction between the RBDs is due to between Lys231 and Asp271 [1].
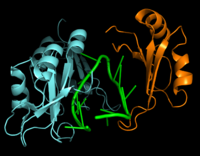
Figure 2: Interaction between Hrp1 (blue), RNA15 (orange) and RNA (green).
Interaction with RNA15
RNA15 is another RNA-binding protein with a single N-terminal RNA recognition motif (RRM) [2]. RNA15 recognizes an A-rich positioning element (PE) downstream from the PEE but upstream from the 3' cleavage site [2]. The recognition of the PE by RNA15 is crucial for precise cleavage of the RNA molecule. Hrp1 and RNA15 are held together by a separate protein, RNA14 [2]. These proteins act together to anchor the polyadenylation and cleavage protein machinery relative to the cleavage site for precise 3'-end processing [2].
Relationship to other proteins

Figure 3: Sequence logo for residues 167-169 of Hrp1. The logo displays the frequency of residues occuring at specific positions within Hrp1. W168 is always conserved in Hrp1 and RRMs of similar proteins.
The RNP-type RBD is found in many proteins involved in post-transcriptional pre-mRNA processing (5'-end capping, splicing, 3'-end cleavage and polyadenylation, and transport from the nucleus)[4]. The unique RBD of Hrp1 enables the protein to bind an RNA sequence that differs in both length and content from the RNA sequences of other RNA-binding and mRNA processing proteins such as sex lethal, Poly (A)-binding protein (PABP), and HuD [1]. Like Hrp1, each of these proteins belong to the class of single strand proteins composed of two canonical RBDs; however, these proteins are differentiated by their target RNA sequence, their interactions with RNA at the atomic level, and their interdomain contacts [1]. One way in which Hrp1 differentiates itself from these other proteins is by the fact that Hud, sex lethal, and PABP all contain at least one intra-RNA base-base stacking interaction, a feature that is not found in the Hrp1-PEE complex [1]. It is possible that the intra-RNA interactions found in these other proteins is replaced by the crucial Trp168-Ade4 stacking interaction found in the Hrp1 complex [1]. The fact that the intra-RNA base-base stacking interactions are replaced by the Trp168-Ade4 in the Hrp1-PEE complex might also explain why the Hrp1-RNA interface involves only 6 nucleotides whereas PABP, sex lethal, and HuD require a longer 8-10 nucleotide sequence in the RNA binding pocket [1].



