Molecular Playground/CsoR and RcnR
From Proteopedia
| |||||||||||
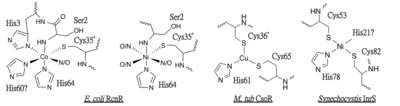
3D structures of copper homeostasis protein
References
- ↑ Reyes-Caballero H, Campanello GC, Giedroc DP. Metalloregulatory proteins: metal selectivity and allosteric switching. Biophys Chem. 2011 Jul;156(2-3):103-14. Epub 2011 Apr 5. PMID:21511390 doi:10.1016/j.bpc.2011.03.010
- ↑ Foster AW, Patterson CJ, Pernil R, Hess CR, Robinson NJ. Cytosolic Ni(II) sensor in cyanobacterium: nickel detection follows nickel affinity across four families of metal sensors. J Biol Chem. 2012 Apr 6;287(15):12142-51. doi: 10.1074/jbc.M111.338301. Epub 2012, Feb 22. PMID:22356910 doi:http://dx.doi.org/10.1074/jbc.M111.338301
- ↑ Iwig JS, Chivers PT. Coordinating intracellular nickel-metal-site structure-function relationships and the NikR and RcnR repressors. Nat Prod Rep. 2010 May;27(5):658-67. Epub 2010 Mar 5. PMID:20442957 doi:10.1039/b906683g
- ↑ Smaldone GT, Helmann JD. CsoR regulates the copper efflux operon copZA in Bacillus subtilis. Microbiology. 2007 Dec;153(Pt 12):4123-8. PMID:18048925 doi:10.1099/mic.0.2007/011742-0
- ↑ Liu T, Ramesh A, Ma Z, Ward SK, Zhang L, George GN, Talaat AM, Sacchettini JC, Giedroc DP. CsoR is a novel Mycobacterium tuberculosis copper-sensing transcriptional regulator. Nat Chem Biol. 2007 Jan;3(1):60-8. Epub 2006 Dec 3. PMID:17143269 doi:http://dx.doi.org/10.1038/nchembio844
- ↑ Foster AW, Patterson CJ, Pernil R, Hess CR, Robinson NJ. Cytosolic Ni(II) sensor in cyanobacterium: nickel detection follows nickel affinity across four families of metal sensors. J Biol Chem. 2012 Apr 6;287(15):12142-51. doi: 10.1074/jbc.M111.338301. Epub 2012, Feb 22. PMID:22356910 doi:http://dx.doi.org/10.1074/jbc.M111.338301
- ↑ Iwig JS, Leitch S, Herbst RW, Maroney MJ, Chivers PT. Ni(II) and Co(II) sensing by Escherichia coli RcnR. J Am Chem Soc. 2008 Jun 18;130(24):7592-606. Epub 2008 May 28. PMID:18505253 doi:10.1021/ja710067d
- ↑ Liu T, Ramesh A, Ma Z, Ward SK, Zhang L, George GN, Talaat AM, Sacchettini JC, Giedroc DP. CsoR is a novel Mycobacterium tuberculosis copper-sensing transcriptional regulator. Nat Chem Biol. 2007 Jan;3(1):60-8. Epub 2006 Dec 3. PMID:17143269 doi:http://dx.doi.org/10.1038/nchembio844
- ↑ Higgins KA, Hu HQ, Chivers PT, Maroney MJ. Effects of select histidine to cysteine mutations on transcriptional regulation by Escherichia coli RcnR. Biochemistry. 2013 Jan 8;52(1):84-97. doi: 10.1021/bi300886q. Epub 2012 Dec 24. PMID:23215580 doi:http://dx.doi.org/10.1021/bi300886q
- ↑ Higgins KA, Chivers PT, Maroney MJ. Role of the N-terminus in determining metal-specific responses in the E. coli Ni- and Co-responsive metalloregulator, RcnR. J Am Chem Soc. 2012 Apr 25;134(16):7081-93. doi: 10.1021/ja300834b. Epub 2012 Apr, 11. PMID:22471551 doi:http://dx.doi.org/10.1021/ja300834b
- ↑ Foster AW, Pernil R, Patterson CJ, Robinson NJ. Metal specificity of cyanobacterial nickel-responsive repressor InrS: cells maintain zinc and copper below the detection threshold for InrS. Mol Microbiol. 2014 May;92(4):797-812. doi: 10.1111/mmi.12594. Epub 2014 Apr 14. PMID:24666373 doi:http://dx.doi.org/10.1111/mmi.12594
Also See
Proteopedia Page Contributors and Editors (what is this?)
Carolyn Carr, Hsin-Ting Huang, Heidi Hu, Alexander Berchansky, Michal Harel