User:Emily Lawson/Sandbox 1
From Proteopedia
Contents |
Poly(A) Binding Protein (PABP)- Homo sapiens
|
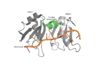
Human Poly(A) Binding Protein (PABP) is a biopolypeptide involved in recognizing the 3'poly (A) tail of mRNA that is added to an mRNA transcript during mRNA processing.This recognition as well as PABP's interaction with other proteins and initiation factors causes it to also play a significant role in translation initiation and mRNA stabilization and degradation. PABP consists of four conserved domains of RNA recognition motifs (RRMs); however, the two N-terminal RRMs (RRM1 and RRM2) and the short linker sequence that connects them supports most of the function of PABP, so RRM3 and RRM4 may not be essential. Thus, the published X-ray structure exhibits RRM1 and RRM2 at a 2.6 Angstrom resolution. This is shown as . Both RRM 1 and 2 are needed to support biochemical function, that is, no one RRM can support biochemical function. Additionally, there is a proline rich C-terminal portion of variable length that is not well conserved and unknown as to how it contributes to the protein's function.
Function
Poly (A)Binding
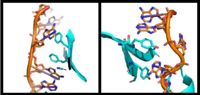
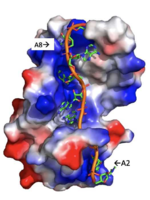
A primary function of PABP is recognizing and interacting with the 3'poly (A) tail created in mRNA processing. As found by EMSA competition experiments, there is a minimum of 11-12 adenosines necessary in the poly (A) tail for the adenosine chain to bind to PABP with high affinity. However, for one biological assembly, a chain containing 9 adenosines sufficiently binds the assembly for crystallization and is shown in the biological assembly structure. The 4 RRM domains that are the primary interacting sites for the adenosine recognition exist as globular domains, each having four antiparallel β-strands and two α-helices. With the N-terminal to C-terminal motifs labeled as S1 to S4 for the β-strands and H1 to H2 for the α-helices, the strands are spatially arranged as S2-S3-S1-S4. Furthermore, there are two conserved sequences in each RRM, called RNP1 and 2. RNP 1 consists of a conserved sequence of 8 residues, while RNP2 consists of a conserved sequence of 6 residues. Much of the weak intermolecular interactions with adenosine from the RRMs occur from the conserved sequences, which correspond to the two central β-strands, with specific interactions shown in Figure 2.The support for adenosine recognition by the RRMs occurs as a type of binding trough with the sheets, primarily , and the interstrand loop between β-strands 2 and 3 as well as the domain linker forming the . Additionally, the primary binding trough is stabilized by .
Adenosine Stabilization Interaction Patterns
Specifically, there are several significant interaction patterns that stabilize adenosine recognition. RRM 1 and 2 makes significant interactions with the adenosine backbone, shown in Figure 3. Additionally, the adenosine stabilizes itself within the binding by intramolecular stacking interactions between adenosines. Through the extensive , the RRM specifies the position of adenosine 2, allowing it to make strong intramolecular stacking interactions with adenosine 1. As a result, adenosine 1 requires less contact with the RRM, as it is mostly stabilized by adenosine 2. Furthermore, some adenosines like adenosine 3 and adenosine 6 are stabilized by being sandwiched between aromatic and alipathic side chains. occurs between aromatic and alipathic side chains and is specified by Lysine 104, and occurs similarly, but it is specified doubly by two residues, Trp-86 and Gln-88.
Translation Initiation
Disease
The initiation of translation in eukaryotes is supported by a closed loop model. This model requires the 5' end and the 3' end of mRNA to be physically connected. The poly(A)-binding protein is necessary for initiation of translation and is required for the closed loop model. Rotavirus, a virus of varying size, containing 11 double stranded RNA and 12 proteins (6 structural, 6 non-structural) is responsible for preventing initiation of translation in infected cells. The virus enters the cell and undergoes a non-conservative replication cycle in the cytoplasm. After a replication cycle non-structural protein 3, NSP3, can be found spread throughout the cytoplasm. NSP3 is responsible for releasing PABP from eIF4F and inhibiting translation initiation. In a study done by Piron et al. it has been seen that NSP3 competes with PABP in binding to the poly(A)-tail of mRNA. This competitor inhibits the proper closing of the closed loop therefore inhibiting translation and protein synthesis. Not only does the rotavirus inhibit protein synthesis of the host cell but it successfully initiatives its own translation as well. The viral mRNA and the host translation initiation factors are in close enough proximity to allow the viral mRNA bound to NSP3 to undergo translation. The translation of viral mRNA allows the virus to spread throughout an organism and lead to a greater decrease in host protein synthesis. When infected with rotavirus one may experience diarrhea, fever, vomiting, and dehydration. Without an antiviral it is suggested to increase fluid intake and allow three to seven days for the infection to subside.
Relevance
Structural highlights
This is a sample scene created with SAT to by Group, and another to make of the protein. You can make your own scenes on SAT starting from scratch or loading and editing one of these sample scenes.
</StructureSection>