User:Juliet Obi/Sandbox 1
From Proteopedia
Contents |
Bromodomain Adjacent to Zinc Finger domain 1A (BAZ1A)
|
Overview
Bromodomain adjacent to zinc finger 1A (BAZ1A) is a protein in humans encoded by the BAZ1A gene. The protein encoded by the BAZ1A gene contains the accessory subunit of the ATP-dependent chromatin assembly factor (ACF), which is a member of the imitation switch (ISWI) family of chromatin remodeling complexes including BAZ1B, BAZ2A and BAZ2B. BAZ1A structure contains a plant homeodomain (PHD) zinc finger at the N-terminus, a bromodomain at the C-terminus, a WAC (Williams Syndrome Transcription Factor WSTF/ ATP-utilizing chromatin assembly and remodeling factor 1 ACF1/ mouse protein related to ACF1 Cbp146) motif found in WSTF/ACF1/cbp146-related proteins [1] and a LH (leucine-rich helical domain) motif [2]. BAZ1A is also known as ATP-utilizing chromatin assembly and remodeling factor 1 (ACF1), Williams syndrome transcription factor-related chromatin remodeling factor 180 (WCRF180), hACF1 or WALp1 (WSTF-, ACF1- like protein 1) [3] [4].
BAZ1A function
BAZ1A encodes the chromatin-remodeling factor ACF1, which is a member of the ISWI chromatin remodeling complexes including the ATP-dependent chromatin assembly factor ACF, and the chromatin accessibility complex, CHRAC. BAZ1A (ACF1) has been implicated in a number of functions including chromatin remodeling, assembly and DNA repair. Together with the ISWI subunit, it has been shown to assemble regularly spaced nucleosome arrays in an ATP-dependent manner [5]. BAZ1A functions in certain DNA repair pathways including nucleotide excision repair (NER), non-homologous end-joining (NHEJ) and homologous recombination (HR), through interaction with a catalytic ATPase subunit, SMARCA5 [6]. BAZ1A has been shown to have a regulatory function in the transcriptional suppression of vitamin D3 receptor-regulated genes [7], and in the transcriptional regulation of stress-induced depressive-like behaviors [8]. A recent study with an identified variant in BAZ1A found that this affects the transcriptional regulatory function of ACF1, thus affecting the expression of genes crucial in vitamin D metabolism, the Wnt signalling pathway, and in proper synaptic function, highlighting the important role BAZ1A has in nervous system development and function [9].
Structural features and comparison with WAL family member BAZ1B
BAZ1A bromodomain histone-ligand recognition
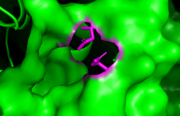
BAZ1A and BAZ1B belong to the WAL family of bromodomains. Both have a conserved asparagine "anchor" residue which in functional bromodomains, hydrogen bonds with the carbonyl group of a bound acetyl-lysine ligand and it is crucial for the interaction. This suggests that BAZ1A and BAZ1B could potentially bind an acety-lysine ligand [10]. Also, BAZ1B has an hydrophobic valine "gatekeeper" residue which is found in the binding pockets of several acetyl-lysine binding bromodomains [11]. Surprisingly, the BAZ1A bromodomain gatekeeper was found to be a non-canonical glutamic acid residue which was found to be compatible with the bromodomain fold through the solving of the crystal structure of BAZ1A bromodomain (indicated in red in the figure below). The glutamic acid residue introduces a negative charge on the side of the supposed binding pocket of BAZ1A which reduces its affinity for acetylated histone ligands [12].Sequence alignment of BAZ1A and BAZ1B bromodomains

BAZ1A-PHD binding to DNA
The PHD finger of bromodomains serves as an epigenetic reader by recognizing the N-terminal histone H3 tail when it is methylated [14]. The function of BAZ1A-PHD, N-terminal to its bromodomain, is however unknown [12]. This can be attributed to the fact that BAZ1A-PHD lacks the "aromatic cage" as well as other key residues crucial for the recognition of trimethylated lysine [15]. BAZ1A-PHD also lacks acidic residues which particularly recognize unmethylated histone H3 lysine 4 (H3K4) peptides [16]. These information show proof that the PHD finger of BAZ1A might interact with DNA through its other biochemical properties other than the canonical residues which are absent [12].
The PHD of BAZ1A is structurally similar to its paralog, BAZ1B-PHD and the dual-zinc binding motif characteristic of their PHD fold have been found to be structurally conserved. BAZ1A-PHD is 54% identical and 73% similar to BAZ1B-PHD [12]. However, BAZ1A-PHD has been found to contain a positively charged feature possibly due to the closely spaced lysine residues on the K1181 and K1183 positions (indicated in red in the image below). The positive charge could be involved in binding a negatively charged partner such as DNA and studies have shown binding affinities tests to confirm this [12]. Although the N-terminal WAC motif of BAZ1A has been shown to bind nucleosomes by a linker DNA in the regulation of the BAZ1A-SMARCA5 complex (ACF complex) [17], the BAZ1A PHD-DNA interaction has been shown not to be needed for this process [12].
Sequence alignment of BAZ1A and BAZ1B PHD fingers

Ligand-interaction domain
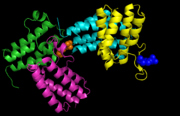
BAZ1A is a monomer and has its structure solved with a unique ligand, 4-(2-hydroxyethyl)-1-piperazine ethanesulfonic acid (EPE), on chains B (1601) and D (1601). A pymol structure showing the ligands in spheres on chains B and D is shown in the image below (PDBID: 5UIY)
BAZ1A protein-ligand interaction
Conservation
BAZ1A appears to be conserved 100% among various species apart from humans. Popular organisms with conserved BAZ1A including humans are mice, zebrafish, rats and bovine animals. BAZ1A also has the evolutionarily conserved glutamic acid gatekeeper which reduces its affinity for binding to acetyl-lysine histone ligands. Substitution of the glutamic acid with a canonical valine gatekeeper has been found to increase its binding to acetyl-lysine histone ligands as well as DNA damage hypersensitivity. This suggests the evolution of BAZ1A to bind acetylated histone ligands with a specific affinity and increasing or decreasing this affinity may prevent BAZ1A from functioning in DNA damage recovery [12].
Disease implications of BAZ1A
There is limited knowledge on the functions of BAZ1A in disease conditions. Recent studies have shown the role of BAZ1A in neurodevelopment and a potential link of BAZ1A to intellectual disability (ID) by showing data of its role on three pathways including vitamin D regulation, Wnt signalling, and postsynaptic signalling [9]. These findings support the growing evidence for the role of chromatin-remodeling factors in cognition with more studies to be carried out in the future.
Medical importance
BAZ1A in complex with SMARCA5 has been implicated in different DNA repair mechanisms [18]. A decrease in BAZ1A expression impairs recovery from DNA damage in human cells [19] [20]. Mutations in the PHD of BAZ1A can disrupt DNA binding and cause transcriptional deregulation before or after DNA damage. The specific target(s) of BAZ1A to DNA damage sites remain unknown, although there is an hypothesis that the DNA binding surface of BAZ1A-PHD is required for disengaging BAZ1A from chromatin after a DNA repair process but it needs to be tested. However, studies have shown the importance of BAZ1A in DNA damage recovery, partly by regulating the fucntion of ISWI chromatin remodelers at DNA damage sites and supporting transcriptional activity required for survival [12].
References
- ↑ Fyodorov DV, Kadonaga JT. Binding of Acf1 to DNA involves a WAC motif and is important for ACF-mediated chromatin assembly. Mol Cell Biol. 2002 Sep;22(18):6344-53. PMID:12192034
- ↑ Jones MH, Hamana N, Nezu Ji, Shimane M. A novel family of bromodomain genes. Genomics. 2000 Jan 1;63(1):40-5. doi: 10.1006/geno.1999.6071. PMID:10662543 doi:http://dx.doi.org/10.1006/geno.1999.6071
- ↑ Poot RA, Dellaire G, Hulsmann BB, Grimaldi MA, Corona DF, Becker PB, Bickmore WA, Varga-Weisz PD. HuCHRAC, a human ISWI chromatin remodelling complex contains hACF1 and two novel histone-fold proteins. EMBO J. 2000 Jul 3;19(13):3377-87. PMID:10880450 doi:http://dx.doi.org/10.1093/emboj/19.13.3377
- ↑ Racki LR, Yang JG, Naber N, Partensky PD, Acevedo A, Purcell TJ, Cooke R, Cheng Y, Narlikar GJ. The chromatin remodeller ACF acts as a dimeric motor to space nucleosomes. Nature. 2009 Dec 24;462(7276):1016-21. doi: 10.1038/nature08621. PMID:20033039 doi:http://dx.doi.org/10.1038/nature08621
- ↑ Ito T, Levenstein ME, Fyodorov DV, Kutach AK, Kobayashi R, Kadonaga JT. ACF consists of two subunits, Acf1 and ISWI, that function cooperatively in the ATP-dependent catalysis of chromatin assembly. Genes Dev. 1999 Jun 15;13(12):1529-39. PMID:10385622
- ↑ Aydin OZ, Vermeulen W, Lans H. ISWI chromatin remodeling complexes in the DNA damage response. Cell Cycle. 2014;13(19):3016-25. doi: 10.4161/15384101.2014.956551. PMID:25486562 doi:http://dx.doi.org/10.4161/15384101.2014.956551
- ↑ Ewing AK, Attner M, Chakravarti D. Novel regulatory role for human Acf1 in transcriptional repression of vitamin D3 receptor-regulated genes. Mol Endocrinol. 2007 Aug;21(8):1791-806. doi: 10.1210/me.2007-0095. Epub 2007 May, 22. PMID:17519354 doi:http://dx.doi.org/10.1210/me.2007-0095
- ↑ Sun H, Damez-Werno DM, Scobie KN, Shao NY, Dias C, Rabkin J, Koo JW, Korb E, Bagot RC, Ahn FH, Cahill ME, Labonte B, Mouzon E, Heller EA, Cates H, Golden SA, Gleason K, Russo SJ, Andrews S, Neve R, Kennedy PJ, Maze I, Dietz DM, Allis CD, Turecki G, Varga-Weisz P, Tamminga C, Shen L, Nestler EJ. ACF chromatin-remodeling complex mediates stress-induced depressive-like behavior. Nat Med. 2015 Oct;21(10):1146-53. doi: 10.1038/nm.3939. Epub 2015 Sep 21. PMID:26390241 doi:http://dx.doi.org/10.1038/nm.3939
- ↑ 9.0 9.1 Zaghlool A, Halvardson J, Zhao JJ, Etemadikhah M, Kalushkova A, Konska K, Jernberg-Wiklund H, Thuresson AC, Feuk L. A Role for the Chromatin-Remodeling Factor BAZ1A in Neurodevelopment. Hum Mutat. 2016 Sep;37(9):964-75. doi: 10.1002/humu.23034. Epub 2016 Jul 8. PMID:27328812 doi:http://dx.doi.org/10.1002/humu.23034
- ↑ Flynn EM, Huang OW, Poy F, Oppikofer M, Bellon SF, Tang Y, Cochran AG. A Subset of Human Bromodomains Recognizes Butyryllysine and Crotonyllysine Histone Peptide Modifications. Structure. 2015 Sep 4. pii: S0969-2126(15)00329-9. doi:, 10.1016/j.str.2015.08.004. PMID:26365797 doi:http://dx.doi.org/10.1016/j.str.2015.08.004
- ↑ Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert JP, Barsyte-Lovejoy D, Felletar I, Volkmer R, Muller S, Pawson T, Gingras AC, Arrowsmith CH, Knapp S. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell. 2012 Mar 30;149(1):214-31. PMID:22464331 doi:10.1016/j.cell.2012.02.013
- ↑ 12.0 12.1 12.2 12.3 12.4 12.5 12.6 12.7 Oppikofer M, Sagolla M, Haley B, Zhang HM, Kummerfeld SK, Sudhamsu J, Flynn EM, Bai T, Zhang J, Ciferri C, Cochran AG. Non-canonical reader modules of BAZ1A promote recovery from DNA damage. Nat Commun. 2017 Oct 11;8(1):862. doi: 10.1038/s41467-017-00866-0. PMID:29021563 doi:http://dx.doi.org/10.1038/s41467-017-00866-0
- ↑ 13.0 13.1 http://tcoffee.crg.cat/apps/tcoffee/do:regular
- ↑ Sanchez R, Zhou MM. The PHD finger: a versatile epigenome reader. Trends Biochem Sci. 2011 Jul;36(7):364-72. doi: 10.1016/j.tibs.2011.03.005. Epub , 2011 Apr 21. PMID:21514168 doi:http://dx.doi.org/10.1016/j.tibs.2011.03.005
- ↑ Li H, Ilin S, Wang W, Duncan EM, Wysocka J, Allis CD, Patel DJ. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature. 2006 Jul 6;442(7098):91-5. Epub 2006 May 21. PMID:16728978 doi:http://dx.doi.org/10.1038/nature04802
- ↑ Lan F, Collins RE, De Cegli R, Alpatov R, Horton JR, Shi X, Gozani O, Cheng X, Shi Y. Recognition of unmethylated histone H3 lysine 4 links BHC80 to LSD1-mediated gene repression. Nature. 2007 Aug 9;448(7154):718-22. PMID:17687328 doi:10.1038/nature06034
- ↑ Hwang WL, Deindl S, Harada BT, Zhuang X. Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA. Nature. 2014 Aug 14;512(7513):213-7. doi: 10.1038/nature13380. Epub 2014 Jun 29. PMID:25043036 doi:http://dx.doi.org/10.1038/nature13380
- ↑ Aydin OZ, Vermeulen W, Lans H. ISWI chromatin remodeling complexes in the DNA damage response. Cell Cycle. 2014;13(19):3016-25. doi: 10.4161/15384101.2014.956551. PMID:25486562 doi:http://dx.doi.org/10.4161/15384101.2014.956551
- ↑ Lan L, Ui A, Nakajima S, Hatakeyama K, Hoshi M, Watanabe R, Janicki SM, Ogiwara H, Kohno T, Kanno S, Yasui A. The ACF1 complex is required for DNA double-strand break repair in human cells. Mol Cell. 2010 Dec 22;40(6):976-87. doi: 10.1016/j.molcel.2010.12.003. PMID:21172662 doi:http://dx.doi.org/10.1016/j.molcel.2010.12.003
- ↑ Klement K, Luijsterburg MS, Pinder JB, Cena CS, Del Nero V, Wintersinger CM, Dellaire G, van Attikum H, Goodarzi AA. Opposing ISWI- and CHD-class chromatin remodeling activities orchestrate heterochromatic DNA repair. J Cell Biol. 2014 Dec 22;207(6):717-33. doi: 10.1083/jcb.201405077. PMID:25533843 doi:http://dx.doi.org/10.1083/jcb.201405077