Introduction
Histones are a family of proteins that condense DNA into chromatin, and is an octamer composed of two of each protein core; H2A, H2B, H3, and H4. Histones are a globular protein, that often have N- or C-terminal tails. These tails can often be subjected to modifications by enzymes. Methylation of histones is one of the four common histone modifications. Methylation is most common on long tails of H3 and H4 due to the tail being able to enter the active site.[1] A histone can be mono-, di-, or tri- methylated. Once the histone is methylated, the DNA goes from tightly bound heterochromatin to loosely packed euchromatin. The euchromatin allows RNA pol II to bind to the DNA and start transcription. [2] Histone methylation is also a major epigenetic marker, which can be passed down from generation to generation. A epigenetic marker affects the way that genes are expressed, and can either activate or repress DNA. Histone methylation is a major epigenetic marker because it has the ability to change heterochromatin to euchromatin, and vise versa. Alterations in markers have been associated with many diseases.[3] Lysine Methyltrasferase SET7/9 (KMT) is an enzyme that methylates the histone 3 lysine 4 (H3K4) and plays a important role in the transcription of DNA in H. sapiens.[4] It is composed of 259 residues and is a monomer containing a SET domain. There is a two-domain architecture containing a conserved anti-parallel β-barrel and an unusual knot-like structure that creates the active site. It also contains a cofactor that plays a role in the active site.The methylation of H3K4 results in transcriptional activation.[4] The specific methylation of H3K4 does not result in a change in charge because it is a nonpolar group being added to the lysine. A change in charge could result in tighter bound heterochromatin.
Structure
SET Domain
The (HKMT) SET7/9 is 366 amino acids long. The overall structure looks like a dimer although it acts as a monomer. The structure is composed of the ΔSET7/9 domain. The ΔSET7/9 consists of the SET domain along with the pre- and post-SET regions.[5][6] The pre- and post-SET regions are adjacent to SET domain and are cysteine rich.[5][6] The pre-SET cysteine region is located near the N-terminal where the post-SET region is located near the C-terminal of the domain.[5][6] These regions are said to play an important role in substrate recognition and enzymatic activity.[5][6]
The SET domain is mostly defined by with the few .[5] form a β-hairpin that sticks out at a right angle to the surface of the enzyme.[3] The following three residues () accommodate a sharp bend in the peptide chain and the end of the protein adopts an .[3] The two most defining features of the SET domain are the C-terminal tyrosine and the knot-like fold. These two components have been recognized to be essential for (SAM) binding and catalysis.[5] [6] [7] The knot-like fold contains the binding sites for the cofactor SAM and the peptide substrate.[8]
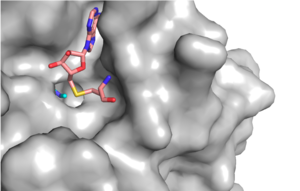
Figure 1: Narrow Lysine Channel
Active Site and Channel
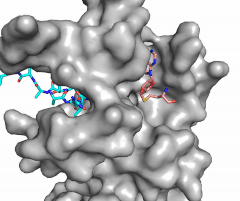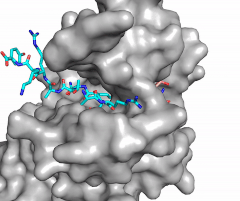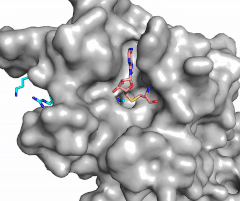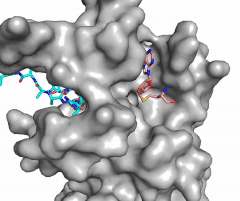
Figure X: Image of substrate bound on one side of SET7/9 (PDB 1o9s) with the lysine target in the active site channel (cyan) and S-adenosyl homocysteine (pink) bound on the opposite face of the enzyme
The most notable feature of the HKMT is the presence of the lysine access channel as the active site. The cofactor and substrate are actually located on opposite sides of the SET domain but are connected through this narrow channel (Figure 1).[3] This channel allows these two components to interact and complete the methyltransfer. The active site in general is considerably tyrosine rich. Residues Tyr245, His297, Ser268, Tyr305, Tyr335, and Tyr337 all help to shape the and the channel.[3] The cofactor involved, SAM, provides the methyl for methylation of the lysine on its sulfur atom.
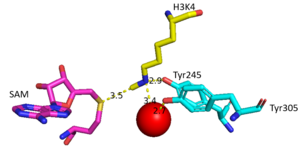
Figure 2: Water being utilized in the active site
The beta hairpin stabilizes the , while also shaping one side of the channel which the peptide binds to.[3] The is composed of residues 255-268.[3] Lysine would have trouble coming down into the active site in its charged form, but it is facilitated by the faces of the flanking tyrosines.[3] The orientation of the lysine is such that the amine-methyl bond is aligned towards the sulfur on SAM so that it can provide the methyl. There is an important water in the active site (Figure 2) as well that acts as a stabilizer for lysine, and helps to shift the lone pair on the nitrogen towards the sulfur of SAM. [3]
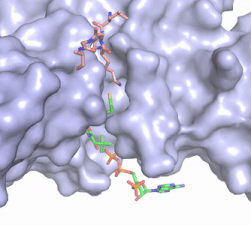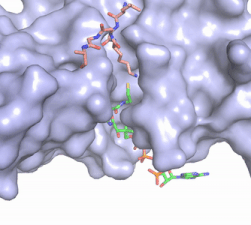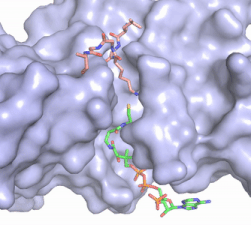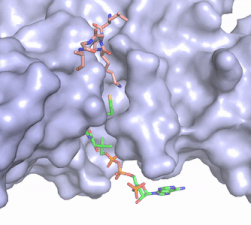
Figure X: The substrate (pink sticks) and Coenzyme A (green sticks) binding channel in Hat1
Function
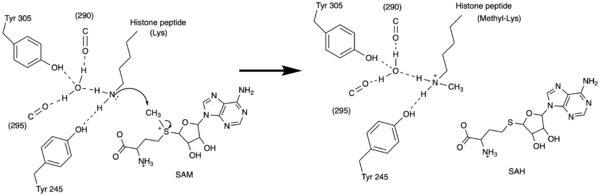
Figure 3: Histone Methylation by HKMT Mechanism
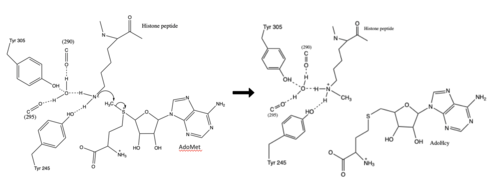
Figure 3: Histone Methylation by HKMT Mechanism
The N from the lysine serves as a nucleophile that attacks the electrophilic CH3 that is present in the AdoMet. The sulfur that the CH3 is attached to pulls the electrons towards itself to weaken the bond between the sulfur and the carbon. This weak bond allows for the N to break that bond and take the methyl group. The N on the lysine is being stabilized by Tyr residues and a water molecule (Figure 3). This allows the N to gain the methyl and take up that positive charge.
The conserved regions of KMT has been found to be vital for function. Multiple studies have found that a mutation to any of the conserved tyrosine residues continues to monomethylate SAM, while it also creates di-methylation and tri- methylation of SAM.[9] In a recent study, Y245A and Y305F were created through site-directed mutagenesis.[9] Due to size, Ala245 was found to create a larger opening of the channel than tyrosine, which allowed for further methylation of SAM. [9] However, Y305F also showed the same characteristics of di- and tri-methylation, most likely due to a decrease of tyrosine residue interaction with water.[9] As there are four invariant conserved tyrosine residues (Tyr305, Tyr245, Tyr335, Tyr337) in the active site, this finding indicates that the function of KMT is dependent on the presence of tyrosine residues in the active site. [9]
Relevance
Renal Fibrosis
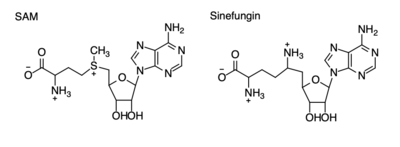
Figure 4: Structural differences between SAM and Sinefungin
The inhibition of KMT SET 7/9 has been found to enhance renal fibrosis.[10] H3K4 methylation activates the transcription of fibrotic genes, and the suppression of the H3K4 methylation was found to enhance renal fibrosis in a mouse model.[10] Sinefungin, a competitive methyltransferase inhibitor, binds to KMT to inhibit SAM.[10] SAM and Sinefungin have similar structures (Figure 4), differing only with the removal of a sulfide and the replacement of a methyl to an amine. The amine replaces the methyl donor for the reaction of KMT, inhibiting the reaction and preventing methylation. Without the methylation of H3K4, the transcription of fibrotic genes is deactivated, leading to renal fibrosis.[10]
Cancer
Many studies have shown that lysine methyltransferases lead to the inhibition of cancers, therefore they are being studied as possible cancer therapy treatments. Lysine methylation contributes to the inactivation of a tumor suppressor gene. Due to this, KMTs are being studied as possible biomarkers for the detection of cancers. Inhibitors of KMT, such as Sinefungin (Figure 4), were used in a study to observe KMT’s effect on cancerous cells. It was found that when KMT is deregulated, tumor behavior increases due to a lack of methylation, indicating the importance of the histone methylation. [11]







