User:R. Jeremy Johnson/RNaseA
From Proteopedia
Contents |
Introduction
Ribonucleases or RNA depolymerases are enzymes that catalyze RNA degradation.[1] Ribonucleases are highly active in ruminants, such as cows, to digest large amounts of RNA produced by microorganisms in the stomach. Ruminants also have high amounts of ribonucleases to process nutrients from cellulose. One such ribonuclease, bovine pancreatic ribonuclease A or RNase A, was one of the most studied enzymes of the 20th century and was used as a model enzyme for many important findings in molecular science. [1] RNase A has been used as a foundational enzyme for the study of protein structure and function, molecular evolution, enzymatic catalysis, protein folding, protein semisynthesis, protein NMR spectroscropy, and protein oligomerization. RNase A is amenable to all of these areas due to its stability, small size, and because the three-dimensional structure is fully determined by its amino acid sequence.[1]
With its importance in molecular science, four researchers have won Nobel Prizes for their work related to RNase A. The 1972 Nobel Prize in Chemistry was awarded to three researchers for their work with RNase A on the folding of chains in RNase A and the stability of RNase A. Christian Anfinsen received the 1972 Nobel Prize in Chemistry for his paper "Principles that govern the folding of protein chains." Stanford Moore and William H. Stein received the 1972 Nobel Prize in Chemistry for their paper "The chemical structures of pancreatic ribonuclease and deoxyribonuclease." The 1984 Nobel Prize in Chemistry was awarded to Robert Bruce Merrifield for his paper "Solid-phase synthesis" using RNase A.[1] RNase A was the first enzyme and third protein for which its amino acid sequence was correctly determined and the third enzyme and fourth protein whose three-dimensional structure was determined by X-ray diffraction analysis [1].[2][3] Disulfide bonds in RNase A were determined after developing a method using Fast Atom Bombardment Mass Spectrometry (FABMS) [2]. The methods of NMR spectroscopy [3] and Fourier transform infrared (FTIR) spectroscopy [4] were developed with RNase A in determining protein structure and protein folding pathways. These new methods, developed with RNase A, could be used for further research to determine the protein structure and protein folding pathways of other proteins.[1]
RNase A and pancreatic ribonucleases have continued to serve as interesting enzymes for study due to the unusual biological actions of ribonuclease homologues. Onconase (ONC)[5] is a structural homologue of RNase A from the oocytes and early embryos of northern leopard frogs. Onconase shows both cytostatic (cell growth suppression) and cytotoxic (prevents cell divisions) characteristics for tumor cells and is currently in clinical trials for the treatment of non-squamous, non-small cell lung cancer.[6] Human pancreatic ribonucleases can be endowed with similar cytotoxic characteristics through specific protein engineering and are undergoing clinical trials for the treatment of late stage solid tumors.[7] Another human homologue of RNase A, angiogenin, is directly involved in neovascularization and mutations in angiogenin have been linked to amyotrophic lateral sclerosis (ALS).[4]
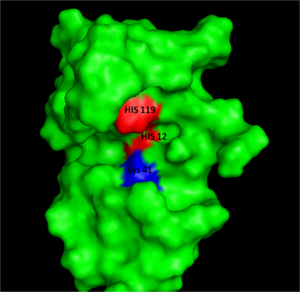
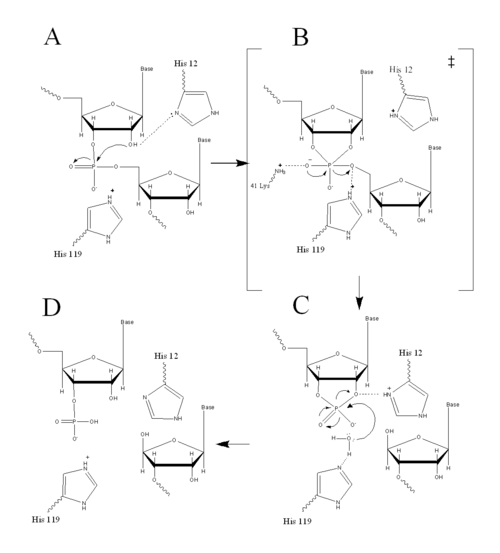
Structure, Catalysis, and Substrate Binding
| |||||||||||
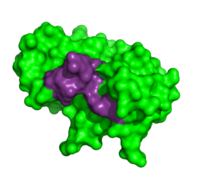
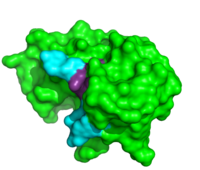
Inhibitors
|
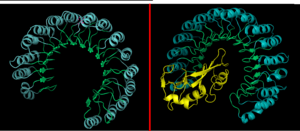
Due to the high rate of RNA hydrolysis by RNase A, mammalian cells have developed a protective inhibitor to prevent pancreatic ribonucleases from degrading cystolic RNA. Ribonuclease Inhibitor (RI) tightly associates to the active site of RNase A due to its . RI is a 50 kD protein that is composed of 16 repeating subunits of alpha helices and beta sheets, giving it a noticeable . The RI-RNase protein-protein interaction has the highest known affinity of any protein-protein interactions with an approximate dissociation constant (Kd) of 5.8 X 10-14 for almost all types of ribonucleases.[11] The ability to be selective for almost all types of RNases, and yet retain such a high Kd is product of its mechanism of inhibition. The interior residues of the horseshoe shaped RI are able to bind to the charged residues of the active site cleft of RNase A, such as . By studying the amphibian RNase, Onconase, the residues Lys7 and Gln11 of RNase A were shown to be the most important in this interaction. In onconase, these residues are replaced with non-charged amino acids, which help prevent the binding of RI to the protein [12]
Additional Proteopedia Pages about RNase A
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 1.9 Raines RT. Ribonuclease A. Chem Rev. 1998 May 7;98(3):1045-1066. PMID:11848924
- ↑ Avey HP, Boles MO, Carlisle CH, Evans SA, Morris SJ, Palmer RA, Woolhouse BA, Shall S. Structure of ribonuclease. Nature. 1967 Feb 11;213(5076):557-62. PMID:6032249
- ↑ Wyckoff HW, Hardman KD, Allewell NM, Inagami T, Johnson LN, Richards FM. The structure of ribonuclease-S at 3.5 A resolution. J Biol Chem. 1967 Sep 10;242(17):3984-8. PMID:6037556
- ↑ Greenway MJ, Andersen PM, Russ C, Ennis S, Cashman S, Donaghy C, Patterson V, Swingler R, Kieran D, Prehn J, Morrison KE, Green A, Acharya KR, Brown RH Jr, Hardiman O. ANG mutations segregate with familial and 'sporadic' amyotrophic lateral sclerosis. Nat Genet. 2006 Apr;38(4):411-3. Epub 2006 Feb 26. PMID:16501576 doi:10.1038/ng1742
- ↑ 5.0 5.1 5.2 'Lehninger A., Nelson D.N, & Cox M.M. (2008) Lehninger Principles of Biochemistry. W. H. Freeman, fifth edition.'
- ↑ 6.0 6.1 6.2 Wlodawer A, Svensson LA, Sjolin L, Gilliland GL. Structure of phosphate-free ribonuclease A refined at 1.26 A. Biochemistry. 1988 Apr 19;27(8):2705-17. PMID:3401445
- ↑ Birdsall DL, McPherson A. Crystal structure disposition of thymidylic acid tetramer in complex with ribonuclease A. J Biol Chem. 1992 Nov 5;267(31):22230-6. PMID:1429575
- ↑ delCardayre SB, Raines RT. Structural determinants of enzymatic processivity. Biochemistry. 1994 May 24;33(20):6031-7. PMID:8193116
- ↑ Thompson JE, Raines RT. Value of general Acid-base catalysis to ribonuclease a. J Am Chem Soc. 1994 Jun;116(12):5467-8. PMID:21391696 doi:10.1021/ja00091a060
- ↑ Fontecilla-Camps JC, de Llorens R, le Du MH, Cuchillo CM. Crystal structure of ribonuclease A.d(ApTpApApG) complex. Direct evidence for extended substrate recognition. J Biol Chem. 1994 Aug 26;269(34):21526-31. PMID:8063789
- ↑ Kobe B, Deisenhofer J. A structural basis of the interactions between leucine-rich repeats and protein ligands. Nature. 1995 Mar 9;374(6518):183-6. PMID:7877692 doi:http://dx.doi.org/10.1038/374183a0
- ↑ Turcotte RF, Raines RT. Interaction of onconase with the human ribonuclease inhibitor protein. Biochem Biophys Res Commun. 2008 Dec 12;377(2):512-4. Epub 2008 Oct 16. PMID:18930025 doi:10.1016/j.bbrc.2008.10.032
External Resources
- Full crystal structure of 1RTA
- Full crystal structure of 1RCN
- Further information on 1RTA
- RNase A- Molecule of the Month
- RNase A-Wikipedia
- Ribonuclease Inhibitor-Wikipedia
- Ribonuclease-Wikipedia
- Ruminants-Wikipedia