[1]
Overview
BRD7 is a human Bromodomain containing protein that is known to be encoded by the BRD7 gene. The full protein has a molecular weight of around 75 kD, while the Bromodomain itself has a molecular weight of roughly 13 kD. It is categorized as a family IV Bromodomain. While its function remains largely unidentified, BRD7 is known to recognize acetylated lysine residues on Histones and play role in regulating transcriptional activity(1). It was first discovered in 2000 in Nasal Pharyngeal Cancer (NPC) cells(2) and has been implicated to have a role in NPC, breast cancer, prostate cancer, and other cancer types (3). BRD7 can recognize acetyl lysine residues by using its Bromodomain to bind to Histone tails. Shown in Figure 1 is a 3D rotating image of the apo BRD7 structure (PDB: 2I7K).
Function
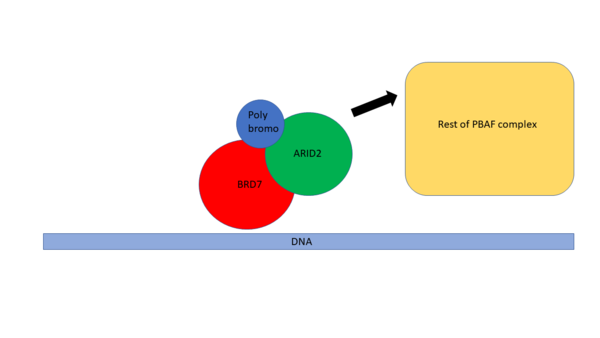
The role of BRD7 in biology is not yet fully understood. It is known however, that BRD7 is part of the PBAF (poly-bromo associated BRG-1 associated factor) complex(4). This is a chromatin remodeling complex that is part of the Swi/Snf family. PBAF is a complex that can facilitate transcription of target genes through chromatin remodeling(5). Shown in Figure 2 is a schematic of BRD7 associating with the PBAF complex. BRD7 was found to be in complexed with Poly bromo (Pb) and ARID2 (6) before associating with the rest of the proteins in the PBAF complex. These proteins can then associate with the PBAF complex to assist in the recognition of post translational marks (PTMs) on Histones.
BRD7 also functions as a tumor suppressor gene that can inhibit p53 and is frequently found to be downregulated in Breast Cancer tumor cells containing wild type p53. BRD7 is important for proper p53 mediated transcription of target genes and is a known p53 co-factor. It likely affects acetylation levels at the promoters of target genes allowing for the facilitation of transcription through regulation of chromatin dynamics(7). BRD7 has also been implicated in having function in maintaining male fertility. Although it is still unclear how, BRD7 is involved in spermatogenesis in male mouse models. Knockout of BRD7 during spermatogenesis resulted in complete infertility(8).
Structural Features
The structure of BRD7 was first solved in 2007 using solution state NMR (9). Like other Bromodomains, BRD7 contains a bundle of 4 left handed antiparallel α-helices. They follow the traditional Bromodomain naming pattern of being called α-Z, α-A, α-B, and α-C in that order from the N-terminal end. In addition to these helices, there are two conserved loops that connect the α-A to the α-Z helix and the α-B to the α-C helix. These are denoted as the ZA and BC loops respectively. To date, two apo structures have been solved for the BRD7 bromodomain using both solution state NMR (9) and X-ray crystallography (10).
Comparison to other family IV bromodomains
BRD has a similar sequence to other Bromodomains contained within family IV. The other proteins in family IV are: BRD9, BRPF3, BRPF1, BRPF1B, BRD1, ATAD2, and ATAD2B. Shown in Figure 3 is a sequence alignment between BRD7 and its closest related member of family IV, BRD9. The bromodomain sequence alignment is 70 residues long for BRD7 and BRD9. For BRD7, the bromodomain sequence ranges from residue 148-218 while in BRD9 the sequence ranges from 153-223. Despite being the two most closely related bromodomains in family IV, these two sequences only share 71.83% sequence identity in the conserved bromodomain region. Amino acid differences with little chemical similarity are highlighted in yellow.
Interactions with Ligands and inhibitors
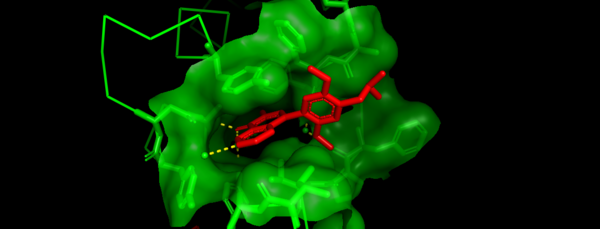
Like other related Bromodomains, BRD7 is also known to bind to acetylated Lysines (Kac) on Histones H3 and H4 (1). Although there is no known structure of BRD7 in complex with an acetylated Histone peptide, there is in vitro evidence that supports the interactions between BRD7 and acetylated Histones (9). Currently the only published structures of BRD7 bound to a ligand has shown it in complex with a variety of well characterized bromodomain inhibitors including BI9564 and BI7273 (9,10). These inhibitors are often competitive inhibitors that will bind the active site of the bromodomain and prevent acetyl lysine binding(11). Shown in Figure 4 is a zoomed in image of BRD7 (green) in complex with bromodomain inhibitor BI9564 (red)(10). BI9564 fits tightly into the acetyl lysine binding pocket of this protein with the main interacting residues being TYR217, ASN211, TYR210, ILE164, PHE158, PHE156, and PHE155(10). Specific mutations in the binding pocket of BRD7 can prevent acetyl lysine recognition as well. Due to its implication in a variety of cancers, and its association with transcriptional regulation BRD7 has become a drug target for therapeutic inhibitors.
Implications in disease
BRD7 along with many other Bromodomains has been widely implicated in a variety of diseases like Cancer(12-14). There is increasing evidence that BRD7 is highly downregulated in a variety of human cancers and BRD7 is known to act as a crucial component of both the p53 and BRCA1 oncogenic pathways(3). BRD7 is most notably associated with NPC. In NPC cells, it is frequently found that BRD7 is under expressed, which may play a large role in NPC development and progression(15). When BRD7 is overexpressed, it slows the growth of NPC cells through transcriptional regulation and inhibits the G1 to S phase cell cycle transition (1).
BRD7 has also been widely studied in breast cancer (13). It is a known binding partner of BRCA1 and has been found to be essential to expression of the Estrogen receptor on breast cancer cells. Inhibition of BRD7 in these cells has been shown to prevent Estrogen receptor expression causing the cancer cells to become more resistant to treatment(13). BRD7 was also found to be frequently deleted or downregulated in breast cancer cells suggesting its importance in the formation or sustainability of these cell lines (12).
References
1. Peng, C., Zhou, J., Liu, H. Y., Zhou, M., Wang, L. L., Zhang, Q. H., Yang, Y. X., Xiong, W., Shen, S. R., Li, X. L., and Li, G. Y. (2006) The transcriptional regulation role of BRD7 by binding to acetylated histone through bromodomain. J Cell Biochem 97, 882-892
2. Staal, A., Enserink, J. M., Stein, J. L., Stein, G. S., and van Wijnen, A. J. (2000) Molecular characterization of celtix-1, a bromodomain protein interacting with the transcription factor interferon regulatory factor 2. J Cell Physiol 185, 269-279
3. Yu, X., Li, Z., and Shen, J. (2016) BRD7: a novel tumor suppressor gene in different cancers. Am J Transl Res 8, 742-748
4. Mashtalir, N., D'Avino, A. R., Michel, B. C., Luo, J., Pan, J., Otto, J. E., Zullow, H. J., McKenzie, Z. M., Kubiak, R. L., St Pierre, R., Valencia, A. M., Poynter, S. J., Cassel, S. H., Ranish, J. A., and Kadoch, C. (2018) Modular Organization and Assembly of SWI/SNF Family Chromatin Remodeling Complexes. Cell 175, 1272-1288 e1220
5. Yan, Z., Cui, K., Murray, D. M., Ling, C., Xue, Y., Gerstein, A., Parsons, R., Zhao, K., and Wang, W. (2005) PBAF chromatin-remodeling complex requires a novel specificity subunit, BAF200, to regulate expression of selective interferon-responsive genes. Genes Dev 19, 1662-1667
6. Kaeser, M. D., Aslanian, A., Dong, M. Q., Yates, J. R., 3rd, and Emerson, B. M. (2008) BRD7, a novel PBAF-specific SWI/SNF subunit, is required for target gene activation and repression in embryonic stem cells. J Biol Chem 283, 32254-32263
7. Drost, J., Mantovani, F., Tocco, F., Elkon, R., Comel, A., Holstege, H., Kerkhoven, R., Jonkers, J., Voorhoeve, P. M., Agami, R., and Del Sal, G. (2010) BRD7 is a candidate tumour suppressor gene required for p53 function. Nat Cell Biol 12, 380-389
8. Wang, H., Zhao, R., Guo, C., Jiang, S., Yang, J., Xu, Y., Liu, Y., Fan, L., Xiong, W., Ma, J., Peng, S., Zeng, Z., Zhou, Y., Li, X., Li, Z., Li, X., Schmitt, D. C., Tan, M., Li, G., and Zhou, M. (2016) Knockout of BRD7 results in impaired spermatogenesis and male infertility. Sci Rep 6, 21776
9. Sun, H., Liu, J., Zhang, J., Shen, W., Huang, H., Xu, C., Dai, H., Wu, J., and Shi, Y. (2007) Solution structure of BRD7 bromodomain and its interaction with acetylated peptides from histone H3 and H4. Biochem Biophys Res Commun 358, 435-441
10. Karim, R. M., Chan, A., Zhu, J. Y., and Schonbrunn, E. (2020) Structural Basis of Inhibitor Selectivity in the BRD7/9 Subfamily of Bromodomains. J Med Chem 63, 3227-3237
11. Clark, P. G., Vieira, L. C., Tallant, C., Fedorov, O., Singleton, D. C., Rogers, C. M., Monteiro, O. P., Bennett, J. M., Baronio, R., Muller, S., Daniels, D. L., Mendez, J., Knapp, S., Brennan, P. E., and Dixon, D. J. (2015) LP99: Discovery and Synthesis of the First Selective BRD7/9 Bromodomain Inhibitor. Angew Chem Int Ed Engl 54, 6217-6221
12. Beroukhim, R., Mermel, C. H., Porter, D., Wei, G., Raychaudhuri, S., Donovan, J., Barretina, J., Boehm, J. S., Dobson, J., Urashima, M., Mc Henry, K. T., Pinchback, R. M., Ligon, A. H., Cho, Y. J., Haery, L., Greulich, H., Reich, M., Winckler, W., Lawrence, M. S., Weir, B. A., Tanaka, K. E., Chiang, D. Y., Bass, A. J., Loo, A., Hoffman, C., Prensner, J., Liefeld, T., Gao, Q., Yecies, D., Signoretti, S., Maher, E., Kaye, F. J., Sasaki, H., Tepper, J. E., Fletcher, J. A., Tabernero, J., Baselga, J., Tsao, M. S., Demichelis, F., Rubin, M. A., Janne, P. A., Daly, M. J., Nucera, C., Levine, R. L., Ebert, B. L., Gabriel, S., Rustgi, A. K., Antonescu, C. R., Ladanyi, M., Letai, A., Garraway, L. A., Loda, M., Beer, D. G., True, L. D., Okamoto, A., Pomeroy, S. L., Singer, S., Golub, T. R., Lander, E. S., Getz, G., Sellers, W. R., and Meyerson, M. (2010) The landscape of somatic copy-number alteration across human cancers. Nature 463, 899-905
13. Harte, M. T., O'Brien, G. J., Ryan, N. M., Gorski, J. J., Savage, K. I., Crawford, N. T., Mullan, P. B., and Harkin, D. P. (2010) BRD7, a subunit of SWI/SNF complexes, binds directly to BRCA1 and regulates BRCA1-dependent transcription. Cancer Res 70, 2538-2547
14. Zhu, B., Tian, J., Zhong, R., Tian, Y., Chen, W., Qian, J., Zou, L., Xiao, M., Shen, N., Yang, H., Lou, J., Qiu, Q., Ke, J., Lu, X., Song, W., Li, H., Liu, L., Wang, L., and Miao, X. (2015) Genetic variants in the SWI/SNF complex and smoking collaborate to modify the risk of pancreatic cancer in a Chinese population. Mol Carcinog 54, 761-768
15. Zhou, J., Ma, J., Zhang, B. C., Li, X. L., Shen, S. R., Zhu, S. G., Xiong, W., Liu, H. Y., Huang, H., Zhou, M., and Li, G. Y. (2004) BRD7, a novel bromodomain gene, inhibits G1-S progression by transcriptionally regulating some important molecules involved in ras/MEK/ERK and Rb/E2F pathways. J Cell Physiol 200, 89-98