User:Sean Callahan/Sandbox 1
From Proteopedia
Lysine Specific Demethylase 1 (Homo sapiens)
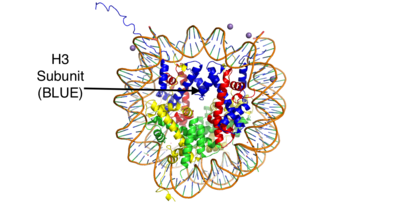
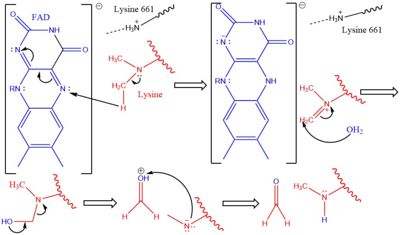
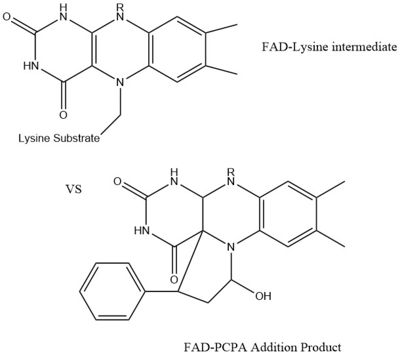
IntroductionHistones are positively charged proteins that help organize DNA into tightly packed chromosomes by acting as a spool for DNA to wrap around. They have an external subunit (H1) that binds linker DNA to string individual histone cores together. The core of a histone is composed of 4 subunits (H2A, H2B, H3, and H4) and has the capability to either loosen their interaction with DNA to promote transcription or tighten their interaction to suppress transcription. Some of the mechanisms that histones use to achieve these different interactions include the addition or removal of acetyl, methyl, or phosphate groups. Demethylases are responsible for removing methyl groups from different histone residues. While this is typically associated with increasing histone-DNA interaction, and thus silencing transcription, demethylation has also been associated with the promotion of transcription depending on the residue that is being demethylated.There are two main classes of demethylases, and they are categorized by their co-factors and co-substrates. One class of demethylases uses an FAD co-factor to catalyze the demethylation reaction. The other class of demethylases uses a Fe2+ ion and α-ketoglutarate as a co-substrate to catalyze the reaction. Although the co-factors used are different, both classes operate by hydroxylating the target methyl group. Lysine Specific Demethylases 1 is a histone demethylase that uses FAD as a co-factor[1]. Specifically, LSD1 is responsible for demethylating Lys 4 and Lys 9 on the H3 subunit of the histone. StructureLSD1 is found in Homo sapiens and its crystal structure was obtained in the presence of the FAD cofactor. LSD1, when compared to other histone binding proteins, contains both conserved and novel structural features. Its substrates are mono- or di-methylated lysine residues and its products are demethylated or mono-methylated lysine residues, respectively. N-TerminusGoing in order of primary structure, the first 166 residues are believed to be unstructured and contain a nuclear localization signal. This area of the protein has also been shown to be susceptible to proteolytic cleavage, which may be to remove the localization signal and render protein inactive[2]. However, a mutant of LSD1, which contains residues 166-852 (essentially eliminating the unstructured region) has been shown to be stable and viable when compared to wild-type LSD1 in a photometric activity assay [2]. Unfortunately, this portion of the protein was unable to be crystallized[2]. SWIRM DomainThe next section of LSD1 spans residues 166-260 and is called the , named after the SWI3, RSC8, and MOIRA proteins from which it was first discovered. It is a conserved domain among histone binding proteins, however LSD1's SWIRM domain is unique in that it does not have a positively charged DNA binding domain on the exterior of the protein. Because of this, it believed that LSD1 does not directly bind DNA unlike other histone binding proteins [3]. The conserved secondary structure of this domain is characterized by a long central helix, with two, shorter helix motifs surrounding it. Oxidase DomainThe is interesting in that it is not completely continuous in the primary structure. The first portion of this domain spans from residues 280-419 and the second portion of the domain spans from residues 520-852, which is the C-terminus of the protein. Its secondary structure is composed of predominantly of alpha helices, with a central 4 turn, anti-parallel beta sheet. This is the largest domain of the protein and houses both the active site and pocket which houses the FAD cofactor. SWIRM-Oxidase InterfaceThe interactions between the SWIRM and Oxidase domains create a cleft right under the . This cleft is made through a number of hydrophobic (van der Waals) interactions. The interior ends of the helices in the SWIRM domain contribute to the cleft, as well as the alpha helices from the oxidase domains. Because of its vicinity to the active site and FAD co-factor, it is believed that this cleft may serve as a site for additional histone tail binding[2]. Tower DomainA unique and defining feature of LSD1 is the 100 residue long insertion between the two parts of the oxidase domain in the primary structure. This spans from residues 419-520. This domain is unique, yet vital to LSD1 function. Specifically, it is hypothesized to be a binding platform of LSD1 to the CoREST complex , as well as a site of allosteric regulation. The CoREST complex is a group of proteins responsible for the silencing of neuronal genes in non-neural cells, and the binding of LSD1 to this complex activates its demethylase activity. It is composed to two long alpha helices(TαA and TαB) that extend from the core of the protein. The helices hold each other in place through hydrophobic interactions. The TαB helix is the shorter of the two and is connected to a helix in the oxidase domain (αD). αD is essential for active site formation, and TαB is thought to be responsible for the correct of αD[2]. RegulationAllosteric StieLSD-1 can be regulated by outside factors. The tower domain and oxidase domain are connected by a , also referred to as the regulator loop in literature. The oxidase domain is what harbors the catalytic chamber for LSD-1 [2]. Any change to the catalytic chamber could drastically reduce the ability for the methylated lysine of the substrate to fit in the binding site correctly. Because the tower domain is attached to the oxidase domain, the tower domain and the connector region may allosterically regulate demethylase activity. Any interaction with these two sites is hypothesized to change the enzyme activity. This has further pushed the assumption that the CoRest complex effects enzyme activity via interaction with the tower domain [2]. Androgen ReceptorLSD-1 will demethylate mono or di-methylated lysines. It will only demethylate lysine 4 of histone 3. This factor can be regulated by the androgen receptor which can change its specificity for a different target. The androgen receptor is a steroid hormone receptor that can modulate transcription. When it interacts with LSD-1 it will no longer demethylate H3-K4, but will now demethylate H3-K9. This attribute allows LSD-1 to work on a wider range of residues. MechanismThe demethylation of lysine via LSD1 occurs through oxidation via a hydride transfer. Shown in Figure 2 the first step involves the initiating a hydride transfer from the one of the two methyl groups bound to the nitrogen at the lysine tail, via a N5 on the flavin group. An imine cation is then formed at the end of the lysine tail to compensate for the loss of hydride. The FAD co-factor is also negatively charged and lysine residue 661 provides stability by drawing that charge away. In the next step the imine is hydrolyzed and transitions to an hemiaminal. This then breaks down to formaldehyde and the product lysine.Hydrophobic PocketThe hydrophobic pocket located in the active site cavity of LSD1, forms a catalytic chamber where the substrate lysine is oriented and positioned to interact with the FAD co-factor to initiate demethylation. The specific residues making up the pocket include valine-317, glycine-330, alanine-331, methionine 332, valine-333, phenylalanine-338, leucine-569, asparagine-660, lysine-661, tryptophan 695, serine 749, serine 760, and tyrosine-761. These residues in the shown in green surrounds the FAD in a way such that it exposes the catalytic nitrogen(N5) that is responsible for the two electron demethylation. The depicted as a sphere is approximately 3.5Å away from the substrate lysine. Lysine 661 shown 4.98Å is responsible for anchoring the FAD in place to efficiently bind the substrate lysine. A structure complex of LSD1 with the a substrate lysine encased in has yet to be crystallized. Another three seperate pockets help bind the histone tail residues to the substrate lysine which is essential for identifying different modifications of the histone tail. ApplicationLSD1 plays a pivital role in many biological processes such as cell growth, epithelial-mesenchymal transition, stem cell biology, malignant transformation of cells, and cell differentiation. Malfunctioning of these processes can lead to life threatening diseases including but not limited to myeloid leukemia and acute lymphoblastic leukemia. Evidence has shown that these activities have ties to prostate and small cell lung cancer so studies have been done to find a reliable inhibitor for LSD1. Monamine oxidases utilize the FAD co-factor much like LSD1 and inhibitors of monamaine oxidase have proven to be successful in inhibiting the activity of LSD1. A patent for 5-cyano indole derivatives(compound similar in structure to monoamine oxidase inhibitors) has been proposed to be a potential inhibitor for LSD1. More specifically trans-2-Phenylcyclopropylamine has been shown the best representation of this inhibition. The formation of a product via an addition reaction through the flavin ring can restrict LSD1's activity with subsequent one‐electron oxidation and cyclopropyl ring opening. However due to the selective nature of 2-PCPA it does not present a strong enough affinity for LSD1's activity. Alternative structures using the foundation of 2-PCPA are continually being researched to find the best fit inhibitor for LSD1. An article published in Wiley Online Library discusses a number of potential LSD1 inhibitors based on monoamine oxidase inhibitors. Thus inhibition of LSD1 opens new pathways for possible treatments for cancers and disorders [4].References
| ||||||||||||