Function
Gene ontology
Molecular function
- 1-acylglycerol-3-phosphate O-acyltransferase activity
- 1-acylglycerophosphocholine O-acyltransferase activity
- O-acyltransferase activity [2]
Biological process
- cardiac muscle contraction
- cardiac muscle tissue development
- cardiolipin acyl-chain remodeling
- cardiolipin biosynthetic process
- cristae formation
- heart development
- hemopoiesis
- inner mitochondrial membrane organization
- mitochondrial ATP synthesis coupled electron transport
- mitochondrial respiratory chain complex I assembly
- muscle contraction
- positive regulation of ATP biosynthetic process
- positive regulation of cardiolipin metabolic process
- regulation of gene expression
- skeletal muscle tissue development [2]
Cardiolipin
Cardiolipin is a trivial name used for 1,3-bis(sn-3’-phosphatidyl)-sn-glycerol. [3] It is a phospholipid mostly found in the inner membrane of mitochondria where is essential for stability of enzymes which are part of the energy metabolism. Cardiolipin is also involved in different stages of the mitochondrial apoptotic process and in mitochondrial membrane dynamics. [4]
Structure
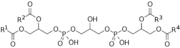
Chemical structure of cardiolipin where R1-R4 are alkyl groups, typically 18-carbon fatty acid side chains.
[5]Cardiolipin’s structure differs from other phospholipids since it is a dimer, it results in a highly specific conical structure. Cardiolipin consists of two phosphatidic acids, three glycerol backbones and four fatty acyl groups, so it can potentially carry two negative charges.[3] [4]
The composition of fatty acids depends on the organism. With a few exceptions like testis or central nervous system, linoleic acid creates 80 % of fatty acids in animal tissues. Testis contain mainly palmitic acid in contrast to central nervous system which contains a wide range of different fatty acids like palmitic, stearic, oleic and many others.[3]
Synthesis
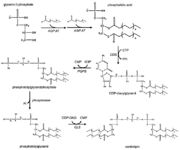
Cardiolipin synthesis.
[6]The first step in the cardiolipin synthesis is a conversion of cytidine diphosphate diacylglycerol to phosphatidylglycerol phosphate by the enzyme phosphatidylglycerol phosphate (PGP) synthase. It is followed by dephosphorylation by PGP phosphatase to phosphatidylglycerol. Lastly the phosphatidylglycerol is converted to cardiolipin in a condensation reaction catalyzed by cardiolipin synthase. [7]
The efficiency of these synthesis depends on the amount of phosphatidyl glycerol in the mitochondrial membrane. The lesser the amount the bigger the effectiveness. [4]
Remodeling
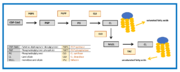
Cardiolipin synthesis and remodeling.
The remodeling is based on unsaturation of the acyl chains of the cardiolipin. This process has two steps. Firstly, the fatty acyl group is removed by a specific cardiolipin deacylase forming monolysocardiolipin. Then tafazzin recycles monolysocardiolipin and makes the final product, unsaturated cardiolipin. [7]
Function
As mentioned, cardiolipin is mainly located on the mitochondrial inner membrane, but is also present on the outer mitochondrial membrane. Cardiolipin is essential for some enzyme systems such as energy metabolism, membrane transport and cell division. For example, when the apoptosis, programmed cell death, or mitophagy, mitochondrial degradation, is needed, cardiolipin is externalized on the surface of mitochondria and acts like a signaling molecule. [3]
On top of that, cardiolipin has many other functions. It is an important cofactor for cholesterol translocation from the outer to the inner mitochondrial membrane. It also has a role in the regulation of gene expression. What is more, it can act like a molecular chaperone, so it helps mitochondrial proteins with folding. [4]
Cardiolipin and Barth Syndrome
When having Barth syndrome, the gene coding enzyme tafazzin is mutated. This is the reason why monolysocardiolipin is accumulated and the unsaturated cardiolipin species are missing. So, in total, patients have a low level of cardiolipin, which is due to its function devastating for the human body. [7]
TAZ gene
Tafazzin is encoded by a TAZ gene (G4.5). This gene is located on the long arm of the X chromosome at position 28 (Xq28). [8]
Several isoforms are known. Isoforms that lack the N-terminus are found in leukocytes and fibroblasts, but not in heart and skeletal muscle. [2]
Human TAZ consists of 11 exons and produces four alternatively spliced mRNA transcripts: a full-length transcript containing all exons (FL), a transcript lacking exon 5 (Δ5), a transcript lacking exon 7 (Δ7) and a transcript lacking both exons 5 and 7 (Δ5Δ7). Of these four alternatively spliced variants of human TAZ, FL and Δ5 encode proteins with transacylase activity, while Δ7 and Δ5Δ7 do not. [9]
Disease - Barth Syndrome
Barth syndrome (BTHS), also known as 3-Methylglutaconic aciduria type II, is an X-linked genetic disorder. The disease is caused by mutation in TAZ gene which encodes for protein tafazzin. [1] Tafazzin works as an acyltransferasein complex lipid metabolism, it is responsible for altering immature cardiolipin- intermediate monolysocardiolipin(with three linoleic acid side chains) (MLCL). [10] [11] Cardiolipin makes up 20% of mitochondrial lipids and is closely connected with the electron transport chain proteins and the inner membrane structure of the mitochondria. [12] Mutations in TAZ gene lead to tafazzin not working properly, immature cardiolipin accumulates whereas the level of cardiolipin is low (mature cardiolopin has four linoleic acid side chains).[10][11] Mitochondria in affected patients are not having a normal shape and functions. Reduced energy production of mitochondria results in apoptosis of cells in tissues with high energy demands, especially cardiac and skeletal muscles. Moreover abnormally shaped mitochondria in white blood cells may affect their ability to proliferate. This causes neutropenia- decreased amount of white blood cells leading to higher risk of infections. [1]
Symptoms
Risk of arrhythmia and sudden death is increased. Neutropenia is most often associated with mouth ulcers, pneumonia, and sepsis. Cardiomyopathy almost always presents before age five. [13]
Diagnosis
Urinary 3-methylglutaconic acid (3-MGC) can be increased 5- to 20-fold, urinary 3-methylglutaric acid and 2-ethylhydracrylic acid may be moderately increased. The diagnosis is established in patient with increased MLCL:CL ratio or detected TAZ pathogenic variant on molecular genetic testing. [13]
3D Structure: Homology Model
No empirical 3D structure for human tafazzin protein (UniProt Q16635) is available in April, 2019. In view of this, homology model was constructed using Swiss-Model server. [14] The closest related protein to tafazzin with a known 3D structure in a public database is in April 2019 plant glycerol 3-phosphate acyltransferase (G3PAT) which has sequence identity of 18,10 %, sequence similarity is 0,29. [14] Tafazzin belongs to a protein superfamily - phospholipid acyltransferases (PF01553).[15]
Domains found on sequence of human tafazzin contain transmembrane domain (15-34) and acyltransferase domain (41-215). [16]

Output from ConSurf analysis of tafazzin showing HX4D motif conservation.

HMM logo of acyltransferase family (PF01553).
[17]In the acyltransferase domain there is a HX4D motif - and residues separated by four less conserved residues. Sequence analysis of membrane-bound glycerolipid acyltransferases revealed that proteins from the bacterial, plant, and animal kingdoms share this highly conserved domain containing .
[18]
Hijikatu, Yura, Ohara and Go in their work from 2015 predicted the intrinsically unstructured regions present in human tafazzin using 15 available prediction servers, eleven of them consistently predicted that the region encoded by exon 5 is intrinsically unstructured, and no other regions of the tafazzin sequence were consistently predicted as intrinsically unstructured.[9]
Hijikatu, Yura, Ohara and Go state in their work of which 2 (Arg94 and Gly197) are replaced more than once. They divided mutated amino acids into 4 groups - amino acids associated with membranes (Asn40, Ile54, Arg57, Arg94, Lys117 - red colored), destabilization (Thr43, Leu50, Gly80, Leu82, Ile209, Leu210, Leu212, His214, Gly216, Gly240 - green colored), substrate binding (Pro62, Ser71, Asp101, Phe104, Ser110, Gly116, Cys118, Val119, Gly124, Phe108, Gly161, Leu169, Trp174, Phe178, Val183, Gly195, Gly197 - blue colored) and catalytic activity (His69 - yellow). [9]



