Molecular Playground/HIV Tat
From Proteopedia
One of the CBI Molecules being studied in the University of Massachusetts Amherst Chemistry-Biology Interface Program at UMass Amherst and on display at the Molecular Playground.
Contents |
Introduction
HIV-1 TAT, or simply Tat, is a human immunodeficiency virus (HIV) gene that regulates transcription of HIV dsRNA.[1] The protein is modeled here to highlight the lack of an alpha helix or a beta sheet. TAT, which stands for trans-activator of transcription, contains 86 amino acid residues in its sequence.[2][3]
|
Cell Penetrating Peptides
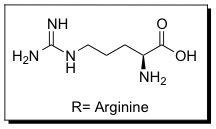
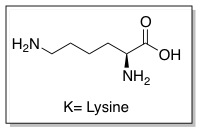
Cell penetrating peptides (CPPs) are proteins with the ability to cross cellular membranes and facilitate the uptake of various cargo, such as small molecules, protiens, antibodies, siRNA, and small DNA fragments.[10] Such cargoes can be associated with CPPs via covalent or non-covalent interactions. HIV Tat is considered a CPP because it contains a protein transduction domain (PTD). PTDs are cation-rich sequences of 10-30 residues, usually containing several Arginine and/or Lysine residues.[11]
These sequences improve interactions between CPPs and cellular membranes and can help them enter cells. The PTD sequence in HIV TAT is YGRKKRRQRRR (amino acid ). The PTD is rich in (Arginine and Lysine ) This is now referred to as the TAT peptide, and has been shown to have improved cellular uptake compared to HIV TAT.
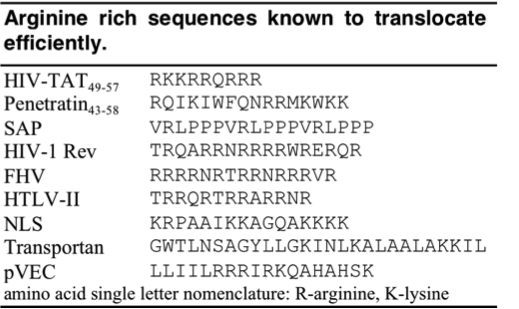
Since the discovery of HIV TAT, other Arginine-rich, natural CPPs have been discovered including, penetratin from Drosophila antennapedia protein , sweet arrow peptide (SAP), HIV-1 Rev, flock house virus (FHV) coat, brome mosaic virus (BMV) Gag, human T-cell lymphotrophic virus (HTLV)-II Rex, and the nuclear localization signal (NLS) from nucleoplasmin. [12][13] The PTD sequences for these proteins can be found in the table below.
TAT-mediated Transduction
The PTD in HIV TAT is highly cationic. When charged residues (Arginine or Lysine) are replaced with neutral residues (Alanine), cellular uptake decreases. It is likely that the cationic charge is necessary for electrostatic interactions with components of the cellular membrane, such as lipid head groups and proteoglycans.[14][15] Other interactions may also play a role in interactions with the cellular membrane, such as hydrophobic and non-covalent interactions.[16] Although cellular uptake is not directly dependent upon architecture (branched vs. linear), it is dependent upon the number of Arginine residues. Polymers with less than five arginine residues are unable to enter cells.[17] As the number of residues increases, cellular uptake improves up until 15 residues where polymers can enter cells but with hindered efficiencies.[18][19] The exact mechanism of cellular uptake is still highly debated in the literature, but a couple of the proposed methods are discussed below.
Transduction
Early studies indicated that HIV TAT primarily entered cells through transduction, meaning that the protein directly crossed the cell membranes. However, for most of these studies, cells were fixed with formaldehyde prior to sample analysis. In 2003, it was discovered that by fixing cells prior to analysis, it permeablized the cell membranes and resulted in artificially high cellular uptake values. Fixed cells showed higher concentrations of TAT in nuclei compared to non-fixed cells where it was localized in the cytoplasm. [20] Studies also showed that FACS could not tell the difference between internalized cells and those proteins that were only bound to the cellular membranes. Cellular uptake studies performed at 4°C and under conditions of ATP depletion show that a small percentage of CPPs can enter cells and also suggests that a majority of the protein enters cells through energy dependent pathways.[21] Some groups have also studied transduction of Tat using model vesicles. [22]
Endocytosis
Another proposed mechanism for cellular uptake is endocytosis, specifically macropinocytosis. This form of fluid phase endocytosis is less known that other types (phagocytosis, clathrin, and coated pit), it is believed to occur in all cell types and enable the uptake of large molecules. [23] During this process, actin protrusions fold around molecules and the surrounding medium and enable cellular uptake.
Tew Group Research Interests
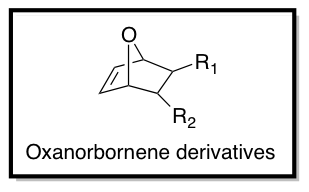
As part of the Tew Research Group in the Department of Polymer Science and Engineering at the University of Massachusetts, Amherst, We are interested in the design of TAT-inspired CPPs for drug and gene delivery. Using functionalized oxanorbornene derivatives as monomers (as shown below), ring opening metathesis polymerization (ROMP) is performed using Grubbs' 3rd generation catalyst to synthesize cationic CPP mimics (CPPMs). Hydrophobicity, charge, aromaticity, and pi-electronics of the monomer structures can be optimized to achieve better cellular uptake and cell viability by changing R1 and R2.
Acknowledgments
The Tew research group is gratefully acknowledged for their contributions to the design and content of this page.
3D structures of Tat protein
References
- ↑ Sodroski et al. Science. 1985. 227, 171-173. [1] http://www.jstor.org/stable/1695050
- ↑ Arya et al. Science 1985. 229, 69-73.
- ↑ Sodroski et al. Science. 1985. 229, 74-77.
- ↑ Dayton et al. Cell. 1986. 44, 941-947.
- ↑ Fisher et al. Nature. 1986. 320, 367-371.
- ↑ Xiao et al. Proc. Natl. Acad. Sci. USA. 2000. 97, 11466-11471.
- ↑ Ensoli et al. Nature. 1990. 345, 84-86.
- ↑ Green et al. Cell. 1988. 55, 1179-1188.
- ↑ Frankel et al. Cell. 1988. 55, 1189-1193.
- ↑ Futaki et al. J. Biol. Chem. 2001. 276, 5836-5840.
- ↑ Wender et al. Adv. Drug Deliv. Rev. 2008. 60, 452-472.
- ↑ Futaki et al. J. Biol. Chem. 2001. 276, 5836-5840.
- ↑ Prochiantz et al. Adv. Drug Deliv. Rev. 2008. 60, 448-451.
- ↑ Ziegler et al. Biochemistry 2003. 42, 9185-9194.
- ↑ Goncalves et al. Biochemistry. 2005. 44, 2692-2702.
- ↑ Ziegler et al. Adv. Drug Deliv. Rev. 2008. 60, 580-597.
- ↑ Wender et al. Proc. Natl. Acad. Sci. 2000. 97, 13003-13008.
- ↑ Mitchell et al. J. Pept. Res. 2000. 56, 318-325.
- ↑ Futaki et al. J. Biol. Chem. 2001. 276, 5836-5840.
- ↑ Richard et al. J. Biol. Chem. 2003. 278, 585-590.
- ↑ Drin et al. J. Biol. Chem. 2003. 276, 31192-31201.
- ↑ Ziegler et al. Biochemistry. 2003. 278, 9185-9194.
- ↑ Gump et al. Trends in Molecular Medicine. 2007. 13, 443-448.