Sandbox Reserved 1236
From Proteopedia
| This Sandbox is Reserved from Jan 17 through June 31, 2017 for use in the course Biochemistry II taught by Jason Telford at the Maryville University, St. Louis, USA. This reservation includes Sandbox Reserved 1225 through Sandbox Reserved 1244. |
To get started:
More help: Help:Editing |
Luciferase
Luciferase is an enzyme responsible for catalyzation of many bioluminencent pathways found in various organism. It is a highly studied and utilized protein for applications in cell biology, genetics and biochemistry.
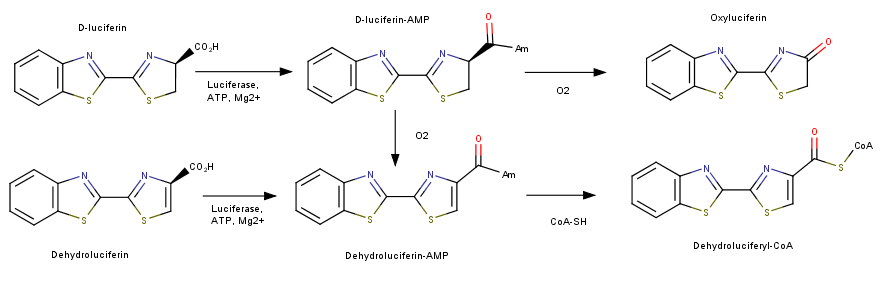
FunctionLuciferase is a class of bioluminescent enzymes that are found in several luminescence organisms. The most studied form Luciferase within its class is found in the North American Firefly (Photinus pyralis). This protein catalyzed the reaction that produces the distinctive yellow flash seen in from the abdomen of the insect. Photinus pyralis is known to use this mechanism for mate attraction and defense. Firefly Luciferase is unique to its species and different forms within the luciferase class can be found in other invertebrates and bacteria.[1]. The most notable being the bioluminescent enzyme found in Click Beetles (Pyrophorus). This protein is one of the most studied and widely used luminescence enzymes having various applications in cell and molecular biology. Luciferase catalyzes two reactions; both reactions use an adenylation reaction along with ATP. The bioluminescent pathway produces Oxyluciferin and a photon of light. The other pathways does not produce light and is known as the nonluminescent reaction. This mechanism utilizes luciferin and produces Luciferyl-CoA. This addiotion of a CoA has been shown to be related to Acyl-CoA synthase used in the activation of fatty acids for oxidation. [2] This image depict the two reactions that luciferase can catalyze. The first being the bioluminencent pathway producing a photon of light and the second, following a mechanism similar to fatty acid synthase, producing a Dehydroluciferyl-CoA.[2] OriginLuciferase is part of the acyl-adenylate/thioester-forming superfamily. Through its two step reaction it ultimately forms an acyl-adenylate intermediate which is esterified into CoA. This mechanism is similar to the other members in this class some of which include Fatty Acyl AMP Ligases, Fatty Acyl CoA Ligases, and Acetyl CoA Ligases. There is evidence that Luciferase has homology with the highly conserved enzymes used in lipid synthesis. [3] Recent studies show that the Gene sequencing of firefly luciferase along with other related preformed and compared to other homologs of the gene. Studies suggest that firefly luciferase gene has similarities to drosophilla fatty acyl-Coa genes. RelevanceThis protein has been utilized in various types of assays ranging from quantification of ATP and the rate of transcription within a cell.[4]. This molecule is especially unique due to the fact that is very efficient in producing a photon through this reaction. Luciferase is sensitive to small changes in substrate and is a optimal choice for quantification of gene expression. It has potential for further biological applications in the future. Luciferase is widely used as a luminescent reporter gene in a variety of assays. The Luciferase gene can be isolated from the firefly and be insterted into a plasmid. This plasmid could contain a gene that is destined for transfection into a cell. After transfection luciferin can be added to a cell culture in order to visualize the expression of the transfected gene. The amount of fluorescence can be quantified to determine the amount of expression within a cell. [5] Structural highlightsThe structure of this protein comprises of The larger one contains an N terminal distorted beta-barrel accompanied by alpha helices. The second and smaller unit is consist of a beta sheet and alpha helix complex [6]. The process of fluorescence is achieved through a two-step oxidation reaction involving the substrate Lucinferin accompanied with ATP, Magnesium and oxygen. The first step consist of using ATP-Mg in an Acylation reaction of the COOH group on Lucinferin producing a Luciferyl adenylate intermediate and a phosphate group. The second reaction uses oxygen to create an excited state of the molecule. The molecule then returns to its ground state emitting a photon of light. [1]
A single peptide has been discover that plays a vital role in the photooxidation by Luciferase. The specific amino acid is a histidine located in the region of the protein [7]. It has been shown to be necessary for the use of oxygen in the second part of the reaction.
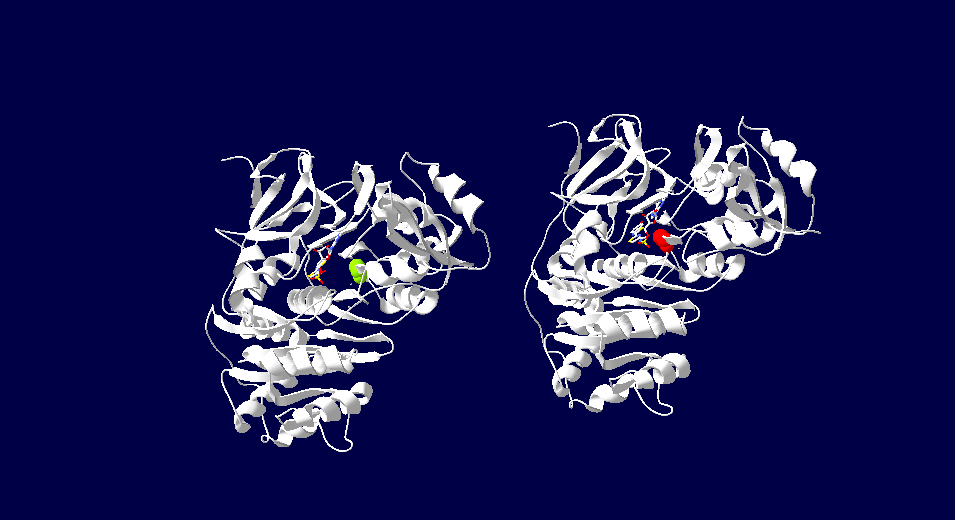
Bioluminescent Structure SpecificationsThe attractive feature for the study and application of luciferase is its bioluminescent activity. The color of light produced has been found to vary depending on the organism that the protein is retrieved from. The fluorsence of light ranges through various wavelengths of light some of which include red, green and yellow light. This difference has a connection to the specificity of the active site of luciferase. Depending on the source of the protein different residues will produce a certain color of fluorescence. Variations in luciferase extracted from japanese fire flies has revealed this specificity. In high energy analogs of luciferase reveal the 286th residue is important in determine the color of light. When comparing green fluorescent japanese luciferin the active site contain an Ser286, while the red emitting variant has an Asp286. These findings indicate that slight variations in the active sight of luciferase can produce various effects on its bioluminencent activity. [8] This image depicts the two forms of Japanese firefly luciferase. The image on the left demonstrates green fluorescense due to its serine286. On the right the luciferase has red fluorescence due to the Asp286. The color of fluorescense is related to the specific amino acids in its active site. [8] InhibitorsInhibition of luciferase can be mediated by several types of molecules. The presence of CoA has been found to have down regulate the bioluminescencent pathway of luciferase. Increased CoA favors the nonluminescent pathway which does not create a photon of light and instead produces Dehydroluciferyl-CoA. In addition to CoA inhibition of this enzyme has been seen with the exposure to diverse group of anesthetics. These molecules bind allosterically to luciferase and cause a conformation change in the protein. This unfolding alters the active site of the protein enough to not allow the bioluminescent catalysis to occur. Other than macro molecules, certain metal ions, specifically Ni and Co, have been seen to inhibit this reaction by replacing Magnesium in the active site. Magnesium is required for use of ATP and without it the reaction will not take place. Of the many inhibitors identified, lipoic acid demonstrates strong activity. Lipoic acid competes with luciferin for the active site enabling the binding of the substrate for the reaction. [9] Luciferase in fungiLuciferase is known to be found in various bacteria and insects, but recent findings suggest that is may also be present in bioluminescent fungi. A similar but unique pathway is currently being studied. With approximately 80 species of bioluminescent fungi known in nature today similar pathways of the luciferase pathway could be discovered in the near future. [10]
References
| ||||||||||||