Introduction
are quinol-dependent transmembrane (Fig. 1) terminal oxidases found exclusively in prokaryotes.[1] With a very high oxygen affinity, bd oxidases play a vital role in the oxidative phosphorylation pathway in both gram-positive and gram-negative bacteria. Cytochrome bd oxidase's responsibility in the oxidative phosphorylation pathway also allows it to act as a key survival factor in the bacterial stress response against antibacterial drugs [1], hypoxia, cyanide, nitric oxide, and H2O2[2]. With their essential roles in bacterial survival, bd oxidases have been pursued as ideal targets for antimicrobial drug development. [3]
![Figure 1. Cartoon model of cytochrome bd-oxidase in E. coli. Dashed lines represent borders of cytoplasmic and periplasmic regions. A quinol bound in the periplasmic jmolSetTarget('1');jmolLink('delete $clickGreenLinkEcho; refresh;setL = \"setLoading();\"; javascript @setL; script /wiki/extensions/Proteopedia/spt/wipeFullLoadButton.spt; ~green = \"Q-loop\"; isosurface DELETE; scn = load(\"/wiki/scripts/83/832924/Q_loop/3.spt\"); scn = scn.replace(\'# initialize;\', \'# initialize;\nclearSceneScaleCmd = \"clearSceneScale();\"; javascript @clearSceneScaleCmd;\n\'); scn = scn.replace(\'_setSelectionState;\', \'_setSelectionState; message Scene_finished;\'); scn = scn.replace(\'_setState;\', \'_setState; setButtonsStartingState();\'); scn = scn.replace(\'DORESIZE\', \'\'); script inline scn;','Q-loop','Q-loop'); is oxidized and releases protons into the periplasmic space, generating a proton gradient. Protons and oxygen atoms from the cytoplasmic side enter cytochrome bd oxidase through specific channels. Oxygen is reduced to water, which is released into the cytoplasmic space. Blue = CydA; green = CydB; yellow = CydX; pink = CydS. [PDB: 6RX4]](/wiki/images/thumb/c/c9/Transmembrane_bd_ox.png/550px-Transmembrane_bd_ox.png)
Figure 1. Cartoon model of cytochrome bd-oxidase in
E. coli. Dashed lines represent borders of
cytoplasmic and
periplasmic regions. A quinol bound in the periplasmic is
oxidized and releases protons into the periplasmic space, generating a
proton gradient. Protons and oxygen atoms from the cytoplasmic side enter cytochrome
bd oxidase through specific channels. Oxygen is
reduced to water, which is released into the cytoplasmic space. Blue = CydA; green = CydB; yellow = CydX; pink = CydS. [
PDB: 6RX4]
The overall mechanism of bd oxidases involves an exergonic reduction of molecular oxygen into water (Fig. 2). During this reaction, a proton gradient is generated in order to assist in the conservation of energy. [4] Unlike other terminal oxidases, bd oxidases do not use a proton pump. Instead, bd oxidases use a form of vectorial chemistry that releases protons from the quinol oxidation into the positive, periplasmic side of the membrane. Protons that are required for the water formation are then consumed from the negative, cytoplasmic side of the membrane, thus creating the proton gradient.
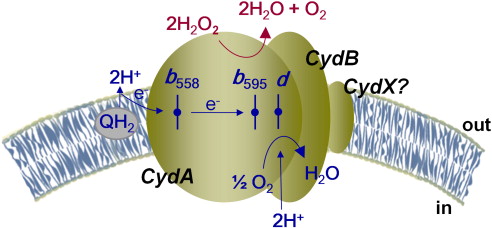
Figure 2. Overall schematic representation of the reductive cycle of cytochrome bd oxidase.
[5] In this cycle, molecular oxygen is reduced into water using the quinol as a reducing substrate. Cytochrome
bd oxidase releases 2 H
+ for each 2 electrons transferred due to the menaquinol oxidation site located on the outer face of the cytoplasmic membrane.
[6] The
bd oxidase completes a redox loop when coupled with quinone
dehydrogenases that receive electrons from
NADH,
pyruvate,
D-lactate, or
acyl coenzyme A. The three hemes essential to the electron transfer are located near the periplasmic space. Heme b
558 is involved in quinol oxidation and heme d serves as the site where O
2 binds and becomes reduced to H
2O. The membrane potential is generated mainly from proton transfer from the cytoplasm towards the active site on the periplasmic side of the membrane.
This page will focus on the structure and overall function of the
bd oxidase in
E. coli. This
bd oxidase is part of the long(L) quinol-binding domain subfamily of terminal oxidases. The L-subfamily of
bd oxidases are responsible for the survival of acute infectious diseases such as
E. coli and
Salmonella. The cytochrome
bd oxidase's three groups, its periplasmically exposed , and will be the primary focus when explaining how the structure of
bd oxidase allows it to catalyze the reduction of molecular oxygen into water.
Structure
Subunits
Cytochrome bd oxidase is made up of four individual subunits.[7] The two major subunits, CydA and CydB, are each composed of one peripheral helix and two bundles of four transmembrane helices. The plays the most important role in the oxygen reduction reaction as it contains the Q-loop as well as all three heme groups. The harbors the molecule which provides structural support to the subunit that mimics the three hemes found in CydA.[1][8] The remaining two subunits, CydS and CydX, are both single helix structures that assist in the oxygen reduction reaction. Unique to E. coli, the binds to CydA to block oxygen from directly binding to heme b595. The promotes the assembly and stability of the oxidase complex. CydX is composed of 37 mostly hydrophilic amino acid residues, including that is exposed to the cytoplasm and prevents the helix from fully entering the membrane. [7]
Q-Loop
Another significant structural feature of bd oxidase is the which is located between TM helices 6 and 7 of the CydA subunit.[7] The periplasmic Q-loop in E. coli stretches over a length of 136 amino acid residues, making it much longer than the Q-loop in Geobacillus Thermodenitrificans.[1] With five helices acting as a flap to cover heme b558, the Q-loop is likely involved in quinol binding and oxidation. The of this Q-loop is very flexible and likely functions as the hinge that allows for quinone binding while the is much more rigid which provides stabilization for the enzyme.[7]
Molecular Function
H and O channels
![Figure 3. H and O-channels of cytochrome bd-oxidase in E. coli. Channels are outlined in gray, water is shown as spheres, and relevant amino acids are labeled above. [PDB:6RX4]](/wiki/images/thumb/8/82/O_AND_H_CHANNEL.png/300px-O_AND_H_CHANNEL.png)
Figure 3. H and O-channels of cytochrome bd-oxidase in
E. coli. Channels are outlined in gray, water is shown as spheres, and relevant amino acids are labeled above. [
PDB:6RX4]
The hydrogen and oxygen channels (Fig. 3) are essential for H+ and O2 molecules to reach the active site of cytochrome bd oxidase. A proton motive force generated by the oxidase[1] allows protons from the cytoplasm to flow through a hydrophilic full of water (pink dots), entering at and moving past [7] where they can be transferred to the active site with the help of the conserved residues [1]. A smaller also exists that transitions from hydrophobic to hydrophilic as it gets closer to the active site. This channel allows oxygen to reach the active site, starting near in CydB and passing by [1], which assists with the binding of oxygen to the active site. The O-channel channel is approximately 1.5 Å in diameter[7], which may help with selectivity.
Interestingly, the O-channel does not exist in the cytochrome bd oxidase of Geobacillus thermodenitrificans; instead, oxygen binds directly to the active site[8]. The subunit found in E. coli blocks this alternate oxygen entry site, which allows oxygen to travel through the O-channel[1][7]. The presence of an O-channel affects oxidase activity, as the E. coli oxidase acts as a "true" oxidase, while the G. thermodenitrificans bd oxidase contributes more to detoxification[7].
Hemes
Three are present in the . These three hemes form a triangle to maximize subunit stability[1][7][8], which is an evolutionary conserved feature across bd oxidases[1]. Heme b558 acts as the primary electron acceptor by catalyzing the oxidation of quinol[7]. Conserved help to stabilize heme b558[7]. Heme b558 transfers the electrons to heme b595, which transfers them to the active site heme d[1]. Multiple residues help stabilzie this electron trasnfer including a conserved that assists heme b595 in transferring electrons to heme d[8]. A conserved is also essential for charge stabilization of heme b595[7], while stabilizes heme d[8]. As heme d collects the electrons from heme b595, in the O-channel facilities the binding of oxygen to heme d, and in the H-channel facilitate proton transfer to heme d[1]. Similar to the three hemes, the (UQ-8) molecule found in the mimics the triangular formation to stabilize the subunit[1].
Mechanism
A reduced quinol with two electrons received from NADH, pyruvate, D-lactate, or acyl coenzyme A transfers these electrons to heme b
558 and releases two protons into the periplasmic space as the initial
electron donor. transfers the electrons to , which transfers the electrons to . Concurrently, the collects protons and the collects oxygen atoms from the cytoplasmic side. The protons and oxygen flow to the active site heme d (Fig. 3). With electrons, oxygen, and protons available, heme d can successfully reduce dioxygen to water (Fig. 2, 4). Using oxygen as the final electron acceptor generates an exergonic reaction that can be coupled with the movement of protons against their gradient when quinol releases two protons into the periplasmic space and when the H-channel uptakes protons from the cytoplasmic side and transfers them to heme d
[1][7].
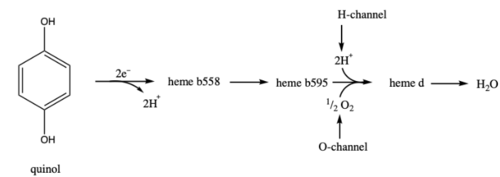
Figure 4. General mechanism of cytochrome bd-oxidase in
E. coli. Electrons are passed from quinol to heme b
558 to heme b
595 to heme d. Protons and oxygen atoms flow into the H-channel and O-channel to heme d. Heme d catalyzes the reduction of oxygen to water.
Relevance
The cytochrome bd oxidase is essential for pathogenic bacteria to thrive in the human body because it enhances bacterial growth and colonization. Any alteration of the bd oxidase Cyd subunits will most likely produce a nonfunctional mutant cytochrome bd oxidase[9], which inhibits bacterial growth. If E. coli are missing or possess ineffective CydA and B subunits, bacterial growth ceases.[10]. With colitis, E. coli mutants that were missing CydAB colonized poorly in comparison to the wild type levels of colonization[10]. The cytochrome bd oxidase is the main component in nitric oxide (NO) tolerance in bacteria, which is released by neutrophils and macrophages when the host is infected[11]. E. coli growth seen in urinary tract infections is mainly due to the NO resistant bd oxidase. Without the CydA and CydB subunits, bacteria could not colonize in high NO conditions[11]. Cytochrome bd oxidases are essential for life in other pathogenic bacteria such as M. tuberculosis. Deletion of the CydA and CydB subunits dramatically decreased the growth of M. tb compared to the wild type when exposed to imidazo[1,2-]pyridine, a known inhibitor of respiratory enzymes[12]. Upregulation of the cytochrome bd oxidase Cyd genes resulted in a mutant strain of M. tb that was resistant to imidazo[1,2-α]pyridine[12].
Since cytochrome bd oxidases are only found in prokaryotes and are required for pathogenic bacterial infections, inhibitors that target cytochrome bd oxidase are promising antibacterial agents. Compounds that target heme b558[2], create unusable forms of oxygen[13], and target the o-channel [14] have shown potential in halting bacterial growth.
![Figure 1. Cartoon model of cytochrome bd-oxidase in E. coli. Dashed lines represent borders of cytoplasmic and periplasmic regions. A quinol bound in the periplasmic jmolSetTarget('1');jmolLink('delete $clickGreenLinkEcho; refresh;setL = \"setLoading();\"; javascript @setL; script /wiki/extensions/Proteopedia/spt/wipeFullLoadButton.spt; ~green = \"Q-loop\"; isosurface DELETE; scn = load(\"/wiki/scripts/83/832924/Q_loop/3.spt\"); scn = scn.replace(\'# initialize;\', \'# initialize;\nclearSceneScaleCmd = \"clearSceneScale();\"; javascript @clearSceneScaleCmd;\n\'); scn = scn.replace(\'_setSelectionState;\', \'_setSelectionState; message Scene_finished;\'); scn = scn.replace(\'_setState;\', \'_setState; setButtonsStartingState();\'); scn = scn.replace(\'DORESIZE\', \'\'); script inline scn;','Q-loop','Q-loop'); is oxidized and releases protons into the periplasmic space, generating a proton gradient. Protons and oxygen atoms from the cytoplasmic side enter cytochrome bd oxidase through specific channels. Oxygen is reduced to water, which is released into the cytoplasmic space. Blue = CydA; green = CydB; yellow = CydX; pink = CydS. [PDB: 6RX4]](/wiki/images/thumb/c/c9/Transmembrane_bd_ox.png/550px-Transmembrane_bd_ox.png)

![Figure 3. H and O-channels of cytochrome bd-oxidase in E. coli. Channels are outlined in gray, water is shown as spheres, and relevant amino acids are labeled above. [PDB:6RX4]](/wiki/images/thumb/8/82/O_AND_H_CHANNEL.png/300px-O_AND_H_CHANNEL.png)

