We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
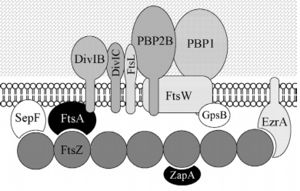
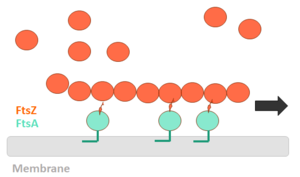
Cell division protein
From Proteopedia
| |||||||||||
References
[10] [11] [12] [13] [14] [15] [16] [17]
- ↑ van den Ent F, Vinkenvleugel TM, Ind A, West P, Veprintsev D, Nanninga N, den Blaauwen T, Lowe J. Structural and mutational analysis of the cell division protein FtsQ. Mol Microbiol. 2008 Apr;68(1):110-23. Epub 2008 Feb 26. PMID:18312270 doi:http://dx.doi.org/10.1111/j.1365-2958.2008.06141.x
- ↑ Alcorlo M, Straume D, Lutkenhaus J, Havarstein LS, Hermoso JA. Structural Characterization of the Essential Cell Division Protein FtsE and Its Interaction with FtsX in Streptococcus pneumoniae. mBio. 2020 Sep 1;11(5). pii: mBio.01488-20. doi: 10.1128/mBio.01488-20. PMID:32873757 doi:http://dx.doi.org/10.1128/mBio.01488-20
- ↑ Angelini S, Deitermann S, Koch HG. FtsY, the bacterial signal-recognition particle receptor, interacts functionally and physically with the SecYEG translocon. EMBO Rep. 2005 May;6(5):476-81. PMID:15815684 doi:10.1038/sj.embor.7400385
- ↑ Galli E, Gerdes K. Spatial resolution of two bacterial cell division proteins: ZapA recruits ZapB to the inner face of the Z-ring. Mol Microbiol. 2010 Jun;76(6):1514-26. PMID:20487275 doi:10.1111/j.1365-2958.2010.07183.x
- ↑ Durand-Heredia J, Rivkin E, Fan G, Morales J, Janakiraman A. Identification of ZapD as a cell division factor that promotes the assembly of FtsZ in Escherichia coli. J Bacteriol. 2012 Jun;194(12):3189-98. doi: 10.1128/JB.00176-12. Epub 2012 Apr, 13. PMID:22505682 doi:http://dx.doi.org/10.1128/JB.00176-12
- ↑ Mosyak L, Zhang Y, Glasfeld E, Haney S, Stahl M, Seehra J, Somers WS. The bacterial cell-division protein ZipA and its interaction with an FtsZ fragment revealed by X-ray crystallography. EMBO J. 2000 Jul 3;19(13):3179-91. PMID:10880432 doi:10.1093/emboj/19.13.3179
- ↑ Duman R, Ishikawa S, Celik I, Strahl H, Ogasawara N, Troc P, Lowe J, Hamoen LW. Structural and genetic analyses reveal the protein SepF as a new membrane anchor for the Z ring. Proc Natl Acad Sci U S A. 2013 Nov 11. PMID:24218584 doi:http://dx.doi.org/10.1073/pnas.1313978110
- ↑ Liu B, Hale CA, Persons L, Phillips-Mason PJ, de Boer PAJ. Roles of the DedD Protein in Escherichia coli Cell Constriction. J Bacteriol. 2019 Mar 26;201(8):e00698-18. PMID:30692172 doi:10.1128/JB.00698-18
- ↑ Silber N, Matos de Opitz CL, Mayer C, Sass P. Cell division protein FtsZ: from structure and mechanism to antibiotic target. Future Microbiol. 2020 Jun;15:801-831. doi: 10.2217/fmb-2019-0348. Epub 2020 Jul , 21. PMID:32692252 doi:http://dx.doi.org/10.2217/fmb-2019-0348
- ↑ Bisson-Filho AW, Discola KF, Castellen P, Blasios V, Martins A, Sforca ML, Garcia W, Zeri AC, Erickson HP, Dessen A, Gueiros-Filho FJ. FtsZ filament capping by MciZ, a developmental regulator of bacterial division. Proc Natl Acad Sci U S A. 2015 Apr 6. pii: 201414242. PMID:25848052 doi:http://dx.doi.org/10.1073/pnas.1414242112
- ↑ Wang, X. & Lutkenhaus, J. FtsZ ring: the eubacterial division apparatus conserved in archaebacteria.Mol. Microbiol. 21, 313–319 (1996). Gueiros-Filho, F. J. & Losick, R. A widely conserved bacterial cell division protein that promotes assembly of the tubulin-like protein FtsZ. Genes Dev. 16, 2544–2556 (2002).
- ↑ Wang, X., Huang, J., Mukherjee, A., Cao, C., and Lutkenhaus, J. (1997). Analysis of the interaction of FtsZ with itself, GTP, and FtsA. J. Bacteriol. 179, 5551–5559.
- ↑ Vaughan S, Wickstead B, Gull K, Addinall SG. Molecular evolution of FtsZ protein sequences encoded within the genomes of archaea, bacteria, and eukaryota. J Mol Evol. 2004 Jan;58(1):19-29. doi: 10.1007/s00239-003-2523-5. PMID:14743312 doi:http://dx.doi.org/10.1007/s00239-003-2523-5
- ↑ Szwedziak P, Wang Q, Bharat TA, Tsim M, Lowe J. Architecture of the ring formed by the tubulin homologue FtsZ in bacterial cell division. Elife. 2014 Dec 9;3:e04601. doi: 10.7554/eLife.04601. PMID:25490152 doi:http://dx.doi.org/10.7554/eLife.04601
- ↑ Huecas, S. et al. Energetics and geometry of FtsZ polymers: nucleated self-assembly of single protofilaments. Biophys. J. 94, 1796–1806 (2008).
- ↑ Lan, G. et al. (2009) Condensation of FtsZ filaments can drive bacterial cell division. Proc. Natl. Acad. Sci. U. S. A. 106, 121–126
- ↑ FILHO, Frederico Gueiros. Cell Division. In: GRAUMANN, Peter L. et al. Bacillus: Cellular and Molecular Biology. Germany: Caister Academic Press, 2017.
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Jonathan Cardoso C. Vieira, Alexander Berchansky, Joel L. Sussman, Jaime Prilusky