Group:SMART:HIV-1 Subtype C Protease
From Proteopedia
|
During the 2009 - 2010 school year, 7 Lincoln students participated in a S.M.A.R.T. Team, under the leadership of Julie Reis of the ALHS Biotechnology Academy. The team worked in partnership with Sabine Jeske & Ben Koo, of UCSF's Science Education Partnership office, and researcher Noah Ollikainen of the Kortemme Lab at UCSF's Mission Bay Campus. S.M.A.R.T. Teams (Students Modeling A Research Topic) were developed by Dr. Tim Herman and Dr. Shannon Colton at the Milwaukee School of Engineering. Lincoln's 2007 team, also mentored by Mrs. Reis, was the first California team to participate in the S.M.A.R.T Team program. S.M.A.R.T. Teams partner with a research scientist to design a three-dimensional model of a particular protein the scientist is studying. The model is designed using a computer program called RasMol and the digital file of the final design is sent to the Milwaukee School of Engineering, where it is translated into a colored, three-dimensional model using rapid prototyping technology.
The Lincoln team met every week after school. The students spent several weeks in the fall semester learning about proteins in general and learning to use the RasMol program. Early in the fall of 2009, the team traveled to the Kortemme lab for a presentation about Noah's research and a tour of the lab. Noah's research focused on HIV-1 Subtype C Protease. The team spent the next few months learning about HIV-1 replication, various current drug regimens for HIV-1 treatment and their mechanisms of action, and prevalence statistics for various strains of HIV-1. In addition to the model of the protease, the team prepared a poster and verbal presentation to explain the enzyme, its role in HIV replication, and the need for more study of HIV-1 subtype C and its protease as a drug target. The group presented its work to researchers at UCSF Mission Bay in March, and then traveled to Anaheim to present their poster and model at the Anaheim Convention Center to scientists attending the 2010 Experimental Biology Conference in April. It was a great year!
Contents |
Abstract
The human immunodeficiency virus 1 (HIV-1) is responsible for the AIDS pandemic that has resulted in the deaths of more than 25 million people since 1981. The HIV-1 viral replication cycle gradually eliminates T helper cells, destroying the immune system and causing death. Once the HIV-1 DNA integrates itself into its host cell’s DNA, the host cell produces immature viral proteins. HIV-1 protease cleaves the immature proteins to form mature HIV-1 viruses. Protease inhibitors prevent the virus from maturing and infecting other cells, and are a common choice for HIV-1 therapy. Due to high mutation rates in all HIV-1 strains, drug resistance is a problem, and for this reason, treatment consists of a combination of inhibitors. HIV-1 subtype C accounts for the highest percentage of infections worldwide and is the most prevalent strain in developing countries. Despite this, most protease inhibitors have been developed to target HIV-1 subtype B, a strain that is most common in Europe and North America.
HIV-1 subtype C protease has two polymorphisms that correlate to resistance to five specific drug combination therapies. A physical 3-dimensional model of HIV-1 subtype C protease may increase understanding how these mutations lead to drug resistance and may result in better drug therapies for patients with subtype C. Our model was built using rapid prototyping at the Center for Biomolecular Modeling at Milwaukee School of Engineering. It is based on the published crystal structure from Coman et al (PDB 2r5p, Biochemistry 2008, 47: 731-743.)
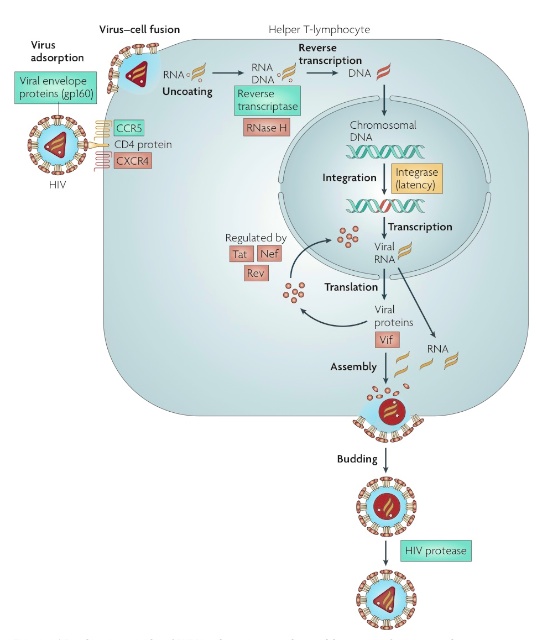
HIV-1 Replication Cycle
HIV binds to a host cell and
releases its capsule into the cell,
revealing two viral RNA strands
and three essential replication
enzymes:
• Reverse transcriptase: used for reverse transcription of viral RNA into DNA.
• Integrase: transfers viral DNA into host cell genome. The host cell transcribes viral DNA into mRNA which leaves to cytoplasm, where immature HIV polypeptides are synthesized.
• HIV protease: cleaves long polypeptides of immature virus into smaller proteins. Two viral RNA strands & essential replication enzymes are bound to immature viral proteins, forming a capsule which then exits host cell to infect other cells throughout the body.
Protease Inhibitors and Drug Resistance
Protease inhibitors block the active site of HIV-1 protease to stop it from cleaving the immature viral polypeptides, preventing viral replication. Due to rapid mutations and the gradual increase in drug resistance, different combinations of drugs must be used. Because subtype B is genetically different from subtype C, the treatments that are used to inhibit HIV subtype B protease are not as effective when used on subtype C. The presence of the mutation M36I on HIV-1 subtype C protease has been shown to correlate with increased drug resistance to several protease inhibitors that are effective against HIV-1 subtype B protease (Indinavir, Atazanavir, Nelfinavir, and Tipranavir).
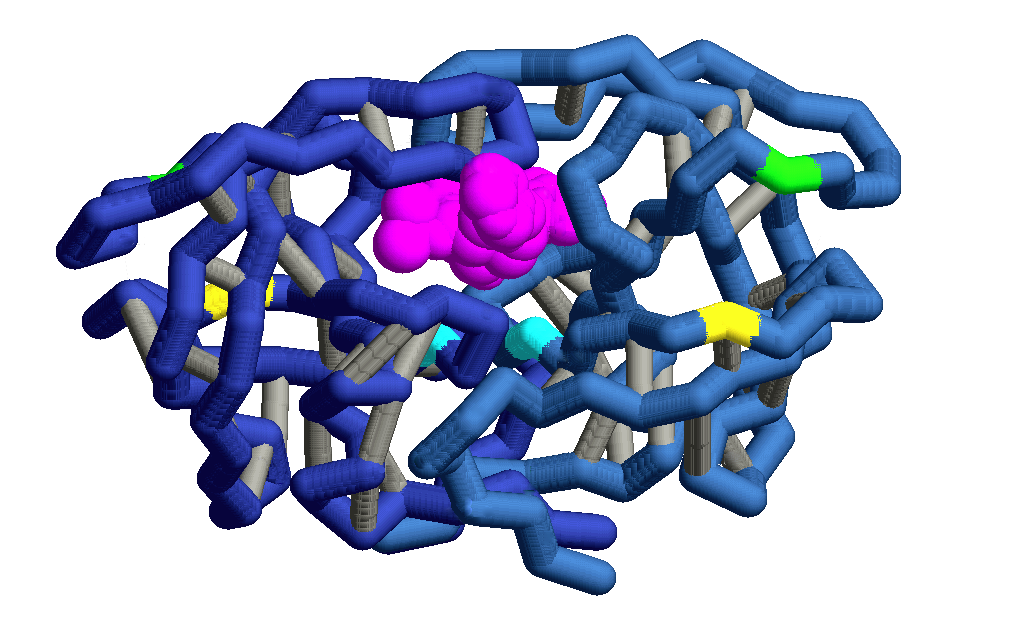
Structural Difference: Subtypes B and C Protease
HIV-1 subtype B and subtype C proteases have very subtle differences in their molecular structures. These subtle differences could render one subtype more prone to becoming resistant to existing drugs than the other subtype. HIV-1 subtype C protease has several amino acid differences that occur outside the active site and have been shown to correlate to drug resistance. These mutations, shown in the model above, are: K20R, M36I, H69K, L89V, and I93L. Although these mutations do not directly affect binding of protease inhibitors, their presence correlates to resistance to several widely used protease inhibitor drugs: it appears that the presence of any one of these mutations may be an initial step towards developing drug resistance. One particular mutation, M36I, is consistently found in HIV strains that are resistant to multiple protease inhibitor drugs,including Atazanavir, Nelfinavir, Tipranavir, and Indinavir. This mutation often occurs along with K20R, which is also associated with resistance to multiple drugs. Both of these mutations, M36I and K20R, have been found in HIV-1 subtype C protease, suggesting that subtype C protease is more prone to developing resistance to these four drugs. A study of the nature of these structural differences may lead to an understanding of the evolution of protease inhibitor drug resistance in HIV-1 subtype C, which in turn may lead to the development of more effective protease inhibiting drugs for this strain.
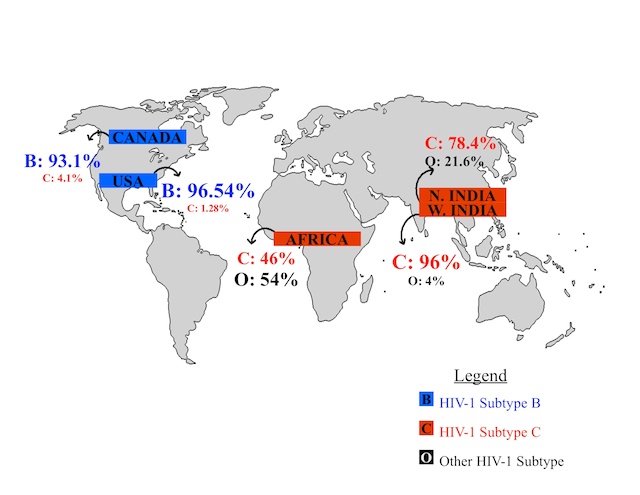
World HIV-1 Subtype B and C Prevalence: What's Wrong With This Picture?
Despite HIV-1 subtype C being the most dominant subtype in Africa and India, which together have the highest incidence of HIV infections and account for over 42% of global HIV infections, HIV research has been focused on HIV-1 subtype B simply due to the prevalence of subtype B infections in the U.S. Since approximately 50% of HIV cases worldwide are attributed to subtype C, the socially responsible and medically sensible thing to do is to direct more research towards understanding the structure of HIV-1 subtype C and developing drugs that are less likely to meet with resistance in subtype C infected patients. The known mutations in HIV-1 subtype C protease offer a logical point of departure for future study, and, ultimately, more successful treatment for these patients.
Additional Resources
For additional information, see: Human Immunodeficiency Virus
References
1. Analysis of HIV Subtype Prevalence and Geographic Distribution in a Large U.S. Population, ARUP Institute for Clinical and Experimental Pathology, Abbott Diagnostics, and University of Utah Department of Pathology.
2. Bessong, P. O.; Polymorphisms in HIV-1 subtype C proteases and the potential impact on protease inhibitors; Tropical Medicine and International Health 13(2) pp. 14-151 (2008)
3. Flexner, C.; HIV drug development: the next 25 years; Nature 6 pp. 959-966 (2007)
4. V. A. Johnson, F. Brun-Vézinet, B. Clotet, H. F. Günthard, D. R. Kritzkes, D. Pillay, J. M. Schapiro, D. D. Richman; Update of the Drug Resistance Mutations in HIV-1: December 2009; International AIDS Society– USA 17(5) pp. 138-145 (2009)