Introduction
Diacylglycerol acyltransferase 1 () is a transmembrane protein that synthesizes triacylglyceridesfrom its two substrates, diacylglycerol (DAG) and fatty acyl-CoA, for dietary fat absorption and fat storage. [1] DGAT1 can be found expressed in the epithelial cells of the small intestine, in the liver where it synthesizes fat for storage, and in the female mammary glands where it produces fat for milk. [1] DGAT1 is a member of the membrane-bound O-acyltransferase (MBOAT) family. [2] All of the enzymes within this MBOAT family are transmembrane enzymes that acylate lipids or proteins. [3] MBOAT enzymes have a conserved MBOAT core, which is a channel-like region that acts as the enzyme’s active site. [3] Additionally, enzymes within the MBOAT family also contain a catalytic histidine within the active site. [3]
Structure
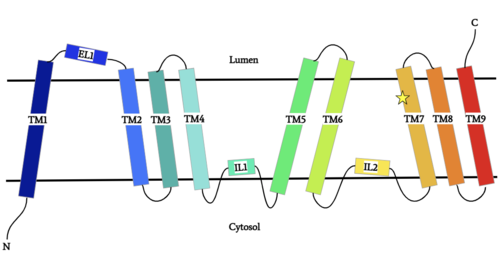
Figure 1: DGAT1 Structure Overview Shown is one transmembrane subunit of DGAT1 with its N-terminus on the cytosolic side and its C-terminus on the luminal side. Labeled are the transmembrane domains (TM 1-7), the intracellular loops (IL1 and IL2), and the ER lumenal loop (EL1). The catalytic Histidine (His415) is labeled with a star on TM7.
Overview
DGAT1 is a dimer that has two identical . Each subunit contains an MBOAT core that acts as its active site. Each subunit also contains (TM), 2 intracellular loops (IL), and one ER lumenal loop (EL).[1] TM2-9, IL1, and IL2 form the structure of the MBOAT core active site. A schematic of DGAT1’s structure is shown in Figure 1.
Dimer Interface
DGAT1 is a dimer that has two identical subunits with 9 transmembrane alpha helices (TM), 2 intracellular loops (IL), and one ER luminal loop (EL). The dimer is held together at the . Both hydrogen bonding and hydrophobic interactions between the residues of the TM1 regions of both subunits act to hold the subunits of the dimer together. [2] Identical domains are bound via measuring 3.19 angstroms between tyrosine 85 and arginine 86 in each subunit. Also present are hydrophobic interactions from the same residues that are formed from pi cloud electrons in the tyrosine ring with the carbon chain in arginine.
Active Site
Overview
Depiction of the DGAT1 active site mechanism is shown in Figure 2. Oleoyl-CoA, the 18 carbon chain derivative of Acyl-CoA, was used to model the substrate Acyl-CoA.[1][2] DAG enters the active site of DGAT1 through its lateral gate while the catalytic His415 flips from the cytosolic side to the lumenal side, in order for the channel opening located on the cytosolic side of DGAT1 to widen enough to accommodate the Oleoyl-CoA.[2] The Oleoyl CoA then enters the active site through the channel opening located on the cytosolic side of DGAT1. When the Oleoyl-CoA and DAG are within close proximity in the active site, the catalytic His415 catalyzes the reaction.[1][2] The coenzyme A moiety exits through the channel located on the cytosolic side, while the product triacylglyceride exits through the lateral gate.
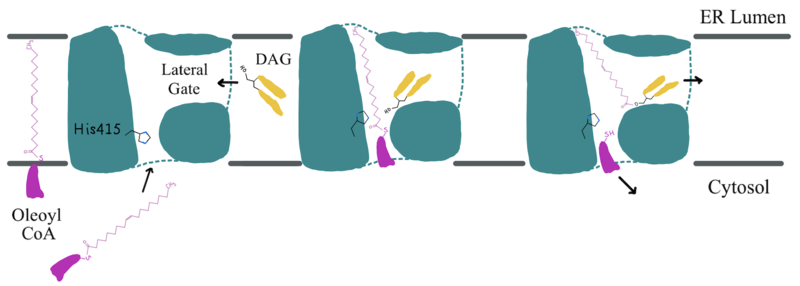
Figure 2: Active Site Mechanism Overview Modeled is the active site of one subunit of DGAT1 (shown in teal) with its catalytic Histidine (His415), Oleoyl CoA (shown in pink), and a general diacylglycerol (DAG, shown in yellow). Shown are the substrates Oleoyl-CoA and DAG entering the active site of DGAT1, being catalyzed by His415, and then leaving through the channel they entered.
Oleoyl-CoA Binding
Oleoyl-CoA enters DGAT1’s active site through a channel on the cytosolic side of the membrane. In order for the channel to widen and accommodate the fatty acid tail of the Oleoyl-CoA, must flip from facing the cytosolic side to facing the lumenal side. It is speculated that the delta sulfur of the Met434 is involved in hydrogen bonding interactions with the catalytic His415 while it is flipped toward the cytosolic side of DGAT1 (6vyi).[2] The His415 needs to break these speculated hydrogen bonds with Met434 to flip toward the lumenal side of DGAT1. Once is bound in the active site, residues Asn378, Gln437, Met434, and Gln465 stabilize the fatty acid tail within the cytosolic channel of the active site, while residues His415 and Gln416 are directly involved within the catalytic mechanism of DGAT1.[2]
DAG Binding
enters the active site through the lateral gate located in the lipid bilayer of the membrane. This lateral gate is a bent and hydrophobic channel that allows for hydrophobic linear or curvilinear molecules to enter.[2] The lateral gate channel is designed to allow for the entrance of DAG and the exit of a triacylglyceride. This channel is also lined with Phe342, Leu261, and Val381.[1][2] Once within the channel, DAG is positioned in close proximity to the bound Acyl-CoA and the catalytic His415.
Active Site Structure
The is located within the pocket formed from TM2-TM9, IL1, and IL2, with the catalytic residue Histidine 415 located on TM7. It is accessible through openings both on the cytosolic and luminal sides. The active site of DGAT1 serves its catalytic function by placing the His415 residue in close proximity to the acyl-CoA in order to cleave its ester bond and bind the fatty acid to the diacylglycerol. The conserved His415 acts catalytically by a charge relay system, where the negative charge of the neighboring Glu416 pulls on the electrons of histidine at the N1 position, making the N3 position more nucleophilic. This nitrogen will then deprotonate DAG so it can begin its attack on Acyl-CoA through acyl substitution. The catalytic mechanism for DGAT1 is shown in Figure 3.[1]
The oxyanion hole of the acyl substitution intermediate should be stabilized by DGAT1. However, the two available PDB files for the substrate-bound DGAT1, 6vz1 and 6vp0, do not show a possible stabilizing residue for the Oleoyl-CoA, the 18 carbon chain version of Acyl-CoA. The PDB file 6vz1 shows the sulfur of Oleoyl-CoA, the electrophile of the mechanism, . This is not likely to be the proper orientation for Oleoyl-CoA, as this sulfur would have to be facing towards the His415 to be protonated after the acyl substitution. For an oxyanion stabilizing residue to be located within this PDB file, the Oleoyl-CoA would have to be properly oriented. The other substrate-bound PDB file 6vp0 shows the sulfur of Oleoyl-CoA facing toward the His415. However, and curved around the oxyanion hole, preventing a stabilizing residue within DGAT1 to be found. For these reasons, an oxyanion stabilizing residue was not identified in the mechanism nor in the images.
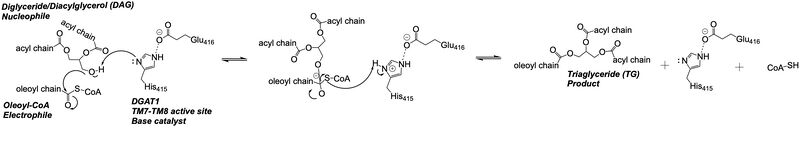
Figure 3: DGAT1 Mechanism The DGAT1 mechanism is an acyl substitution with DAG as the nucleophile and Acyl-CoA as the electrophile. His415 is the catalytic residue that deprotonates DAG to make it a better nucleophile. Glu416 stabilizes the His415. The residue that stabilizes the oxyanion hole of the intermediate is unknown.
Medical Relevance
Disease
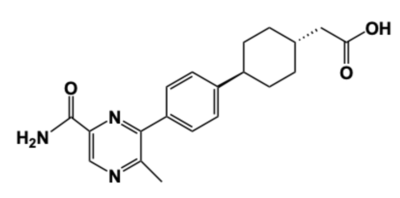
Figure 4: DGAT1 Inhibitor AZD7687 Shown is the structure of AZD7687, a known inhibitor of DGAT1.
Obesity and nonalcoholic fatty liver disease (NAFLD) result from an accumulation of triacylglycerides within the body. Recently, DGAT1 has become a therapeutic target for obesity and nonalcoholic fatty liver disease in order to reduce triacylglyceride storage within the body.[4][5] Different inhibitors have been created, such as AstraZeneca’s direct inhibitor AZD7687, shown in Figure 4.[4] AZD7687 has an EC50 value 0.44 µmol/L, showing that it binds with high affinity at a low concentration of DGAT1.[4] However, while triacylglyceride accumulation decreased, negative side effects did occur, such as diarrhea and other adverse GI symptoms.[4]
Congenital Protein-Losing Enteropathy
Additionally, congenital protein-losing enteropathy (PLE) is linked to DGAT1 mutations. PLE is a GI disorder that causes malabsorption of fat and a deficiency in fat-soluble vitamins. Patients in a congenital PLE case study exhibited a homozygous missense Leu295Pro mutation within the MBOAT core of their DGAT1 enzymes.[6] is located within the MBOAT core active site on TM5. While the Leu295 is not near the catalytic residues His415 and Glu416, the mutation will disrupt the overall active site. Proline is an alpha helix breaker because it causes steric hindrance within the backbone of the helix turn. It is hypothesized that this mutation breaks this helix in the MBOAT core and greatly reduces its enzymatic activity and ability to make triacylglycerides. Without proper DGAT1 function to produce triacylglycerides, there is a decrease in albumin, which is a protein that helps prevent fluid from leaking out of the liver and blood vessels.[5] This decrease in albumin then leads to decreased efficiency in nutrient transport and fat absorption.
Relevance
DGAT1 is an important enzyme in the synthesis of triacylglycerides and has relevance in the research of diseases that involve triacylglyceride accumulation, like obesity and NAFLD, or triacylglyceride reduction, like congenital PLE. There is an opportunity within pharmaceuticals for developing DGAT1 inhibitors like AZD7687 to lessen the severity of various diseases through a decrease in triacylglyceride storage. However, inhibitors will have to work around the symptoms of triacylglyceride reduction, as seen through the impact of the Leu295Pro mutation in congenital PLE patients. Developing an inhibitor that can balance DGAT1 efficacy between excess and deprivation of triacylglyceride synthesis can improve treatments for obesity, NAFLD, and other triacylglyceride storage diseases.




