metabotropic Glutamate Receptor 5
Introduction
G-coupled protein receptors (GPCR's) are helical trans-membrane proteins that bind to an extracellular signal and activate a cellular response. The human genome encodes for approximately 750 GPCR's, 350 of which are known to respond to extracellular ligands.[1] GPCR's are divided into four major classes based on sequence similarity and transduction mechanism: Class A,B,C, and F.[2] Metabotropic Glutamate Receptor 5 () is a class C GPCR that is involved in the Gq pathway.[3] In this pathway, glutamate binds to the extracellular domain of mGlu5, and the trans-membrane domain then undergo a conformational change that activates the coupled G-protein on the intracellular side of the membrane.[4] The activated G-protein disassociates, and the alpha subunit activates Phospholipase C. Phospholipase C in turn cleaves PIP2 to DAG and IP3. IP3 then binds to calcium channels on the Endoplasmic reticulum, creating an increased cellular concentration of calcium. Increased calcium concentrations thus lead to increased neuronal activity.[2] Due to its involvement in neuronal activity, mGlu5 is highly expressed in neuronal and glial cells in the central nervous system, where glutamate serves as the major neurotransmitter.
Structure
Overall Stucture
is seen as a homodimer in vivo, with each subunit being comprised of three domains: extracellular, trans-membrane and cysteine-rich. mGlu5 is centered on the trans-membrane domain, comprised of seven α-helices all roughly parallel.[4] Also displayed is Intracellular Loop (ICL) 1 which forms a short α-helix and interacts directly with a trans-membrane helix to stabilize mGlu5’s conformation. Additionally, ICL3 and Extracellular Loops (ECL) 1 and 3 all lack secondary structure. and ECL2 interacts with trans-membrane (TM) helices 1, 2, and 3 as well as ECL 1, again helping to stabilize the protein’s overall conformation.[4]
Key Interactions
A number of intramolecular interactions within the trans-membrane domain stabilize the inactive conformation of mGlu5, as demonstrated by being represented in the inactivate state. While in the inactive state, glutamate binding to mGlu5 triggers a conformational change that leads to mGlu5 being in the active state and hence initiates the aforementioned Gq pathway. The first of these interactions is an ionic interaction, termed the , between Lysine 665 of TM3 and Glutamate 770 of TM6. Evidence for the importance of this interaction came through a kinetic study of mutant proteins where both residues were separately substituted with alanine, resulting in constitutive activity of the GPCR and its coupled pathway.[4] A second critical interaction that stabilizes the inactive conformer is a between Serine 614 of ICL1 and Arginine 668 of TM3. Similarly, when Serine 614 was substituted with alanine, high levels of activity were seen in the mutant GPCR.[4] A between Cysteine 644 of TM3 and Cysteine 733 of is critical at anchoring ECL2 and is highly conserved across Class C GPCR’s.[4] The ECL2's presence combined with the helical bundle of the trans-membrane domain creates a that restricts entrance to the allosteric binding site within the seven trans-membrane α-helices. This restricted entrance has no effect on the natural ligand, glutamate, as it binds to the extracellular domain, but this entrance dictates potential drug targets that act through allosteric modulation.[4]
Clinical Relevance
Role in Diseases
mGlu5 is located mainly post-synaptically and is in high abundance in the nucleus accumbens, caudate nucleus, striatum, hippocampus and cerebellar cortex.[5] These areas of the brain are highly involved in cognition, motivation, and emotion, which are essential neural functions. Diseases and other mental deficiencies arise from either an overactivation of the GPCR, which overactivates its coupled signaling pathway, or from underactivation of both the GPCR and its coupled pathway.[6] Negative allosteric modulators (NAMs) work to decrease protein activity and are being studied as treatments for fragile X-syndrome, depression, anxiety and dyskinesia.[4] Conversely, positive allosteric modulators (PAMs) work to increase protein activity and are being studied for the treatment of schizophrenia and cognitive disorders.[6]
Interactions with a Negative Allosteric Modulator
The structure of mGlu5 bound to the NAM Mavoglurant demonstrates how protein activity is decreased through drug interactions.
binds within the allosteric binding site in the core of the seven trans-membrane α-helices, having passed through the restricted entrance formed by the . Bound Mavoglurant forms multiple interactions with the protein that further stabilize the inactive conformation.
The bicyclic ring system of the drug is surrounded by a pocket of mainly hydrophobic residues including Val 806, Met 802, Phe 788, Trp 785, Leu 744, Ile 651, Pro 655, and Asn 747 (Figure 1).[4] The carbamate tail of Mavoglurant forms a hydrogen bond through its carbonyl oxygen to the amide side-chain of Asn 747 of TM4. A hydroxyl group similarly forms hydrogen bonds to mGlu5, specifically at two serine residues (S805 and S809) of TM7 (Figure 2). These residues form a hydrogen bonding network to other residues through their main chain atoms and a coordinated water molecule. The interactions between Mavoglurant and mGlu5 involve TM helices that were not previously stabilized by any strong interactions, introducing a new level of stability that favors the inactive conformation of the protein and hence decreases the overall activity of mGlu5.[4]
Mavoglurant was met by disappointing Fragile X-syndrome trial results and ultimately discontinued for Fragile X-syndrome in 2014, but testing for dyskinesia and Obsessive-compulsive disorder are still in progress. Basimglurant and MTEP are inhibitors that are similar to Mavoglurant as they both function as negative allosteric modulators to mGlu5, and they are currently under trials to serve as medications for depression.[7][8]
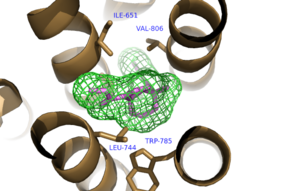
Figure 1. Hydrophobic Pocket Surrounding Mavoglurant
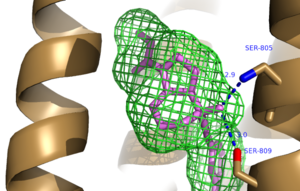
Figure 2. Hydrogen Bonding between mGlu
5 and Mavoglurant. Blue coloration represents Hydrogen Bond donor whereas red coloration represents Hydrogen Bond acceptor.


