Introduction
Histone Methylation
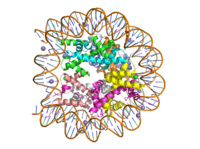
Figure 1: Human nucleosome particle, pbd code: 5y0c
Histone proteins aid in the packing of DNA for the purpose of compacting the genome in the nucleus of the cell and regulating physical accessibility of genes for transcription. The nucleosome (Figure 1) itself is an octamer complex of the core histone proteins H2a, H2b, H3, and H4, which organize into two heterodimers. Additionally H1 and H5 serve as linker proteins. About 145-157 DNA base pairs wind around a histone heterodimer core.[1] Modifications to histone core proteins can affect the accessibility of transcription factors to the genome, either promoting or inhibiting transcription. Some of these modifications include methylation/demethylation, acetylation/deacetylation, and ubiquitination/deubiquitination.[2]
Histone methylation has been found to be associated with both gene activation and deactivation. The factors influencing the specific outcome are the methylation site, residue type (Lys or Arg) and the extent (mono-, di- or tri-methylation).[3] Sites known for gene activation are Lys-4, Lys-36, and Lys-79 on H3; whereas, methylation at Lys-9 and Lys-27 on H3 and Lys-20 on H4 are known for gene inactivation.[4] Typically, methylation of some of these sites are can be found on both active and inactive genes. Some tumor related genes such as p53 are site specifically methylated to promote biological function [4], whereas hypomethylation of CpG is linked to tumor genesis.[2]
Sidechain methylation of basic amino acids in histone proteins is accomplished by the Histone Methyltransferase class of enzymes. In all but one instance, the many enzymes that are lysine substrate specific contain the structural protein domain known as the SET domain. Generally, these are known to methylate lysine in the H3, H4 and H2a core proteins. One enzyme in this family is the , which adds a methyl group to Lys4 of the H3 protein. This histone modification results in promotion of gene unwinding and gene transcription.[5], [3]
Lysine Methyltransferase (KMT) Structure
The structure of human histone methyltransferase SET7/9 was determined by x-ray diffraction at 1.75Å, along with a bound 10 residue peptide representing the H3 substrate protein.[5] SET7/9 was crystallized with its cofactor S-adenosyl methionine (SAM) in its unmethylated form, S-adenosyl homocysteine (SAH), and with the H3 peptide model in its product form, specifically the lysine at residue 4 methylated.
Overall Structure
The SET7/9 enzyme structure sequentially consists of a N-terminal domain (177-193), followed by the characteristic SET domain () which itself ends with a specific C-terminal segment (344-366). The enzyme is best characterized as having alpha+beta folding topology as it consists of a mixture of both alpha helix and beta sheet, but without any significant repeating pattern [5]. The helical composition includes 3 , with two residing in the SET domain and one in the C-terminal segment. The alpha helices in the SET domain are of length two turns while the C-terminal helix is by far the largest with four turns. There are also two in the SET domain which are each one turn. There are 21 total found in both the N-terminal and the SET domains. The beta strands are primarily anti-parallel and multiple antiparallel strands are connected by Type 1 and Type 2 .
The Active Site
The active site and binding pocket of KMT contain residues and shape that optimize catalytic function and stability. First, the lysine of the substrate histone enters the active site via the comprised of Tyr335 and Tyr337. With the histone substrate bound, the alkyl part of the lysine sidechain is stabilized by the , and polar residues are stabilized by hydrogen bonding interactions on the surface. The Tyr335 and Tyr337 are also essential for stabilization of histone chain via hydrogen bonding. The itself contains the cofactor S-adenosyl methionine (SAM) which donates the methyl group in the reaction.[5] In the active site scene, the structures depict the post-reaction result, where the Lys has been methylated and SAM has been converted to S-adenosyl homocysteine (SAH).
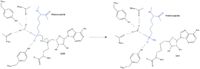
The overall methylation reaction (Figure 2) is stabilized by several hydrogen bond interactions between active site residues and the substrate Lys amine. Specifically, the Tyr305 hydroxyl and the mainchain carbonyl oxygens of Ala295 and Ser290 coordinate with a buried water molecule that in turn coordinates with a free hydrogen off the Lys amine nitrogen. Additionally, the hydroxyl of Tyr245 hydrogen bonds to the other Lys amine hydrogen. These interactions force the amine nitrogen to become more nucleophilic so that subsequently attack of the SAM methyl group carbon becomes favorable. Attack is further facilitated as the cofactor methyl is bound to a positively charged sulfur. Finally, the methyl group is transferred to the amine, the charge on the sulfur is resolved and thus SAM is converted to SAH.[5]
The C-Terminal Domain
The C-terminal domain of lysine methyltransferase is very important for the catalytic activity of the enzyme. The overall structure of the provides various interactions that help stabilize the in the correct orientation for a reaction in the active site.[5] Hydrophobic interactions in the C-terminal domain are mainly responsible for stabilizing the access channel to the methylation site for the H3 lysine. Residues 337-349 create a that stabilizes the orientation of two tyrosine residues Tyr 335 and Tyr337 that form the lysine access channel. Furthermore, the hydrophobic packing of alpha-helix 3 against beta-sheet 19, specifically , stabilize the orientation of the so that its methyl donating group is oriented toward the lysine access channel. The orientation of the SAM cofactor is further stabilized by interactions with C-terminal domain residues .[5]
The N-terminal Domain

Figure 3: Clustal alignment of N-terminals
Though a highly conserved region (Figure 3), the of SET7/9 is notably far from the active site and has not been shown to be involved in enzyme activity or participate in substrate binding.[6] With deletion of the N-terminal domain, studies have shown this modification does not affect SET7/9 activity.[5] Though not essential for catalytic activity, the N-terminal domain may interact with other small molecules or proteins to act as an allosteric regulator region to the C-terminal domain.[6]
Inhibitors
SET7/9’s structure and function has been studied extensively because of its role in transcription [7]. In the past few years it has been identified to methylate genes involved in multiple diseases; making it a potential candidate for drug inhibition.[8] Two compounds that have been found to inhibit SET7/9 in certain cells in vitro are Sinefungin and Cyproheptadine. Each inhibitor acts on the catalytic center of SET7/9, however their mechanisms of inhibition and possible medical relevancies differ greatly.
Sinefungin
Sinefungin is a potent methyltransferase inhibitor that is a natural nucleoside isolated from the "Streptomyces" species.[8] Also referred to as adenosyl-ornithine, it is the delta (5’ adenosyl) derivative of ornithine and a structural analog of SAM. Sinefugin is unique because it binds in the cofactor pocket rather than where the substrate binds like a typical competitive inhibitor (Figure 4). The amine group of sinefungin potentially makes two hydrogen bonds to the main chain carbonyls of Arg265 and His293. As a result, sinefungin is potentially more stable bound in the active site than SAH due to the formation of these two additional hydrogen bonds, which are not possible with SAH’s sulfur.
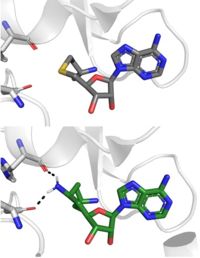
Figure 4: SAH (grey) and Sinefungin (green) in the SET7/9 cofactor binding pocket. The amine group of sinefungin potentially makes two hydrogen bonds to the main chain carbonyls of Arg265 and His293. Sinefungin was modeled using PDB: 1O9S by changing the sulfur of the bound SAH to methylene and then attaching an amine group.
In some renal disease patients using peritoneal dialysis a condition called peritoneal fibrosis develops which reduces how effective fluid is removed.[9] Sinefungin has been used to inhibit the SET7/9 enzyme to treat peritoneal fibrosis in mice and tested against human peritoneal mesothelial cells.[8] SET7/9 is involved in peritoneal fibrosis because it mono-methylates H3K4, which activates the transcription of fibrosis related genes. The administration of Sinefungin to mice in vitro resulted in decreased levels of mono-methylated H3K4 protein (H3K4me1), as well as suppressed peritoneal cell density and thickening. This decrease suggests that the methylation of H3K4 was inhibited by Sinefungin, and furthermore that inhibiting SET7/9 might have the potential to ameliorate peritoneal fibrosis.
Cyproheptadine
Another inhibitor of SET7/9 is cyproheptadine, a clinically-approved anti-allergy drug that was originally developed as histamine (H1) and serotonin (5-HT2A) receptor antagonist.[7] The structure was determined by X-ray diffraction at 2.0 Å resolution with cofactor SAM and with cyproheptadine. Unlike Sinefungin, it is a traditional inhibitor and competitive with the histone peptide substrate as it binds to the peptide-binding site. When cyproheptadine binds to the substrate site, the methylated nitrogen of the piperdine ring forms a hydrogen bond with Thr286 as well as hydrophobic interactions with the residues surrounding its binding site. The binding of cyproheptadine to the catalytic site causes conformational changes of residue Tyr337, an important residue for the formation of the lysine access channel. This movement subsequently causes a conformational change of the . The conformational change ultimately generates a large hole adjacent to the lysine access channel, as well as shift the position of the C-terminal helix.[7]
With the revelation of its inhibitory effects on SET7/9, cyproheptadine was used in vitro to treat MCF-7 breast cancer cells. SET7/9's non-histone activities include the methylation of the estrogen receptor α (ERα), a nuclear receptor and a transcription factor responsible for estrogen-responsive gene regulation. The expression and transcriptional activity of ERα is involved in the carcinogenesis of 70% of breast cancers, making it a major target for hormone therapy. Researchers found that treating the MCF7 cells with cyproheptadine decreased ERα's expression and transcriptional activity which therefore inhibited the estrogen-dependent cell growth. These findings suggest that cyproheptadine could possibly be repurposed to breast cancer therapy in the future.[7]