Publication Abstract from PubMed
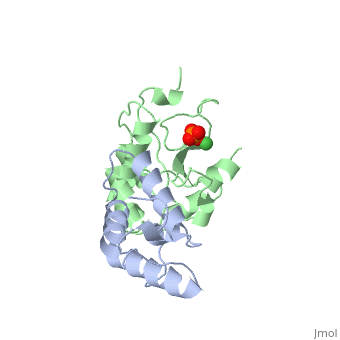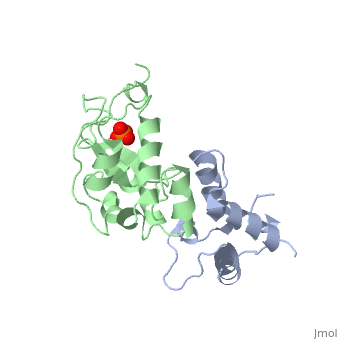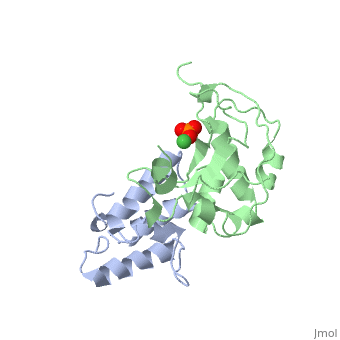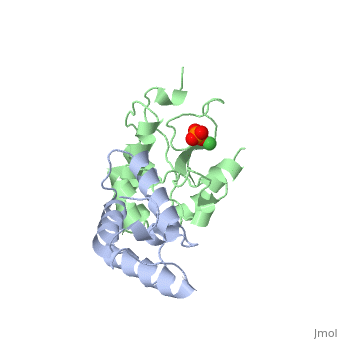
The crystal structure of the cytotoxic endonuclease domain from the bacterial toxin colicin E9 in complex with its cognate immunity protein Im9 reveals that the inhibitor does not bind at the active site, the core of which comprises the HNH motif found in intron-encoded homing endonucleases, but rather at an adjacent position leaving the active site exposed yet unable to bind DNA because of steric and electrostatic clashes with incoming substrate. Although its mode of action is unorthodox, Im9 is a remarkably effective inhibitor since it folds within milliseconds and then associates with its target endonuclease at the rate of diffusion to form an inactive complex with sub-femtomolar binding affinity. This hyperefficient mechanism of inhibition could be well suited to other toxic enzyme systems, particularly where the substrate is a polymer extending beyond the boundaries of the active site.
Structural and mechanistic basis of immunity toward endonuclease colicins., Kleanthous C, Kuhlmann UC, Pommer AJ, Ferguson N, Radford SE, Moore GR, James R, Hemmings AM, Nat Struct Biol. 1999 Mar;6(3):243-52. PMID:010074943
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.
of the immunity protein 9 (Im9, 1bxi, colored yellow), evolved variant R12-2 (lime), and immunity protein 7 (Im7, colored blue, 7cei) reveals their structural identity. However, when the immunity proteins-bound , they demonstrate somewhat different picture. The Im9 and Im7 are differ more in their binding configurations (19°, with Tyr54-Tyr55 as the pivot), while the variant R12-2 is in an intermediate configuration between Im9 and Im7. Of note, in the variant R12-2 (3gkl) and Im9 (1bxi) there are Tyr54 and Tyr55, while in the Im7 (7cei) Tyr55 and Tyr56 are homologous to them. The most are in the loop between helices α1 and α2 in Im9 (yellow, labeled in black) and evolved variant R12-2 (lime, labeled in black). This loop consists of three mutations: N24D, T27A, and S28T in variant R12-2. We can see the deviations in the relative position of helices α1 and α2, in the loop's backbone and in the side chains of residues 24, 26 and 28.
Comparison of the different Im-colicin complexes reveals changes in the binding configuration of the evolved variants which increase affinity toward ColE7 by re-aligning pre-existing Im9 residues. Glu30 of Im9 (1bxi, colored yellow) forms with Arg54 of ColE9 (orange), whereas Asp51 have not direct side chain–side chain interactions. Asp31 of Im7 (blue) which is corresponding to Im9 Glu30 is involved in to Arg520 and Lys525 of ColE7 (darkmagenta), while Asp52 of Im7 (corresponding to Im9 Asp51) is within hydrogen bond distance to Thr531 and Arg530 of ColE7 (7cei). Glu30 in the variant R12-2 (lime) is shifted and forms a to Arg520 of ColE7 (magenta). Asp51 is within hydrogen bond distance to Thr531 of ColE7 (3gkl). However, the side chains of Lys525 and Arg530, which are very important in salt bridge contacts with Glu30 and Asp51, respectively, in the structure of the ColE7–Im7 complex have a different conformation that eliminates these contacts in evolved variant R12-2.