Expression of concern about:
Structural Basis of Enzymatic Activity for the Ferulic Acid Decarboxylase (FADase) from Enterobacter sp. Px6-4
Wen Gu, Jinkui Yang, Zhiyong Lou, Lianming Liang, Yuna Sun, Jingwen Huang, Xuemei Li, Yi Cao, Zhaohui Meng, Ke-Qin Zhang [1]
by Harry M. Greenblatt and Joel L. Sussman
Molecular Tour
Gu et al. [1] reported the crystal structure of ferulic acid decarboxylase (FADase) from Enterobacter sp. Px6-4. The enzyme catalyzes the decarboxylation of ferulic acid, to give 4-vinylguaiacol, a potential precursor for vanillin. The crystals obtained contained two enzyme molecules in the asymmetric unit. One structure was of the “native” conformation, with no substrate bound in the active site of either monomer. In this crystal structure appears to have undergone significant structural changes, with electron density appearing for a . The , and no unusual electron density was observed in the active site. The authors modeled an unreacted ferulate molecule into the density in the active site of the first monomer, and attributed the lack of binding in the second monomer to close crystal contacts, which limit the necessary movements required to allow binding. This latter structure was deposited in the PDB with accession code 3nx2.
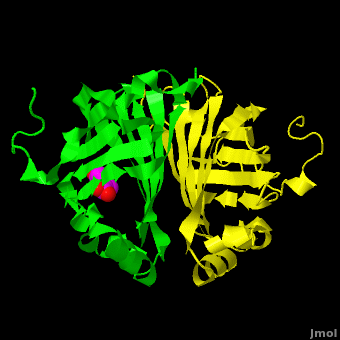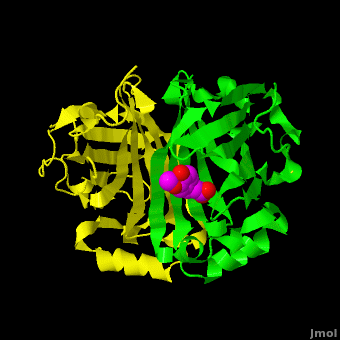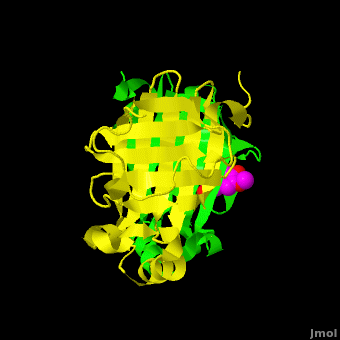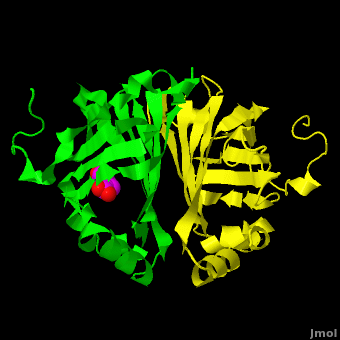
The editors of the journal were contacted by a reader, who raised concerns about the accuracy of the assignment of the density in the of 3nx2. The reader suggested that the density was more reminiscent of a (used in the crystallization buffer) based on the shape of the density, and the fact that the stereochemistry in the region of the double bond (between atoms C7 and C8) was distorted. O atoms are colored in red, N atoms are in blue, S atom in yellow; C atoms of HEPES in cyan, and C atoms of Ferulate in magenta. . In response to these concerns the Editors contacted one of us (JLS), asking for a critical review of the claims of the authors regarding 3nx2. The results of this review are detailed below.
The structure of 3nx2 was downloaded from the PDB as well as the structure factors. The software suite PHENIX[2] was used to run simulated annealing on the structure, with the ligand removed. This produced a simulated annealing Fo - Fc omit map, which should have all structural bias caused by the original ligand removed. A HEPES molecule was fitted into this density using COOT[3], and refined in PHENIX (R = 19.4%, Rfree = 24.4%). Although the R and Rfree are slightly higher than for 3nx2 (R =18.9%, Rfree = 23.6%), this is to be expected, since the geometry of the model refined in PHENIX (rms bonds 0.007, angles 1.12) was much tighter than the one obtained from REFMAC[4] (rms bonds 0.019, angles 1.81). In Figure (see below), we superimpose the omit map with the two refined structures: on the left, 3nx2, with magenta carbon atoms, and on the right, with cyan carbon atoms, the PHENIX-refined enzyme-HEPES complex.
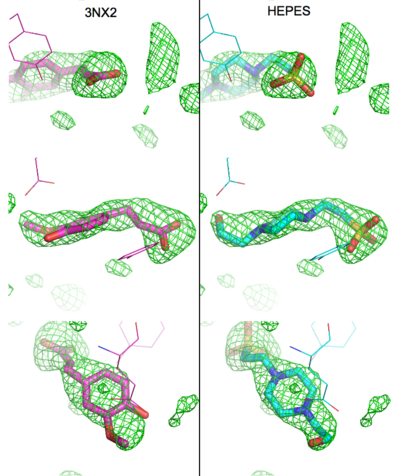
Figure: Simulated annealing omit map (Fo – Fc) in region of active site, contoured at 3σ. On the left is the superposition of the structure of ferulate from
3nx2 (
magenta C atoms), and on the right the refined HEPES molecule (
cyan C atoms). The three views show different aspects of the density and modeled molecules. In addition we have included Tyr27 and Glu134 from each respective structure. Prepared with
PyMOL.
Although ferulic acid does somewhat match the difference density, the HEPES molecule does appear a better match for the electron density, both in terms of the larger sulfone "head" and the ethoxy tail attached to the ring. While these considerations alone may be insufficient to positively identify the observed density as a HEPES molecule, other considerations make the assignment of this density as ferulate rather difficult. As pointed out by the readers, the of ferulic acid should conjugate with the through the of ferulic acid. As such, the (e.g. in the PDB entry 2bjh). Indeed, in examples of this type of compound in the Cambridge Structural Database (small molecules) these systems are all planar. In the PDB there are 12 submissions (excluding 3nx2) with ferulic acid/ferulate bound. All but two have this system as planar. One of the exceptions is a 1.0Å structure, where there is some deviation of the carboxylate group from planarity, and the other 2wtm, where one of the two molecules of ferulate has been modeled with some deviation from planarity. The authors of 2wtm do not discuss this, and looking at the density maps, this appears to be an oversight on their part, since a planar structure would fit their maps. In the of 3nx2, however, a planar molecule would not fit the density well.
Furthermore, it is unclear why the authors feel that sodium ferulate would be an inhibitor. As shown by the authors in Figure 6 of Gu et al. [1], the substrate is ferulate. In fact, the authors themselves, in the second paragraph of the Discussion (page 6) refer to the "...enzyme-substrate complex...". Thus, the soaking experiment simply introduced substrate into the crystal, which underwent decarboxylation. Trapping unreacted substrate in the active site would be rather unlikely, and would require supporting evidence to indicate that this has happened.
In light of the above, we propose the following explanation for the observations in the native structure (3nx1), and in the complex (3nx2).
1. HEPES is a relatively weak competitive inhibitor of FADase. In the native crystals, the binding constant of HEPES was not large enough to effect the conformational change necessary to allow binding in the crystal structure, despite the fact that HEPES was present in the crystallization mixture.
2. In the crystals of the complex, the authors added the substrate, which underwent decarboxylation, and was able to transform the active site of chain A to the bound form. This allowed binding of HEPES to chain A. Crystal contacts kept the active site of chain B locked in the unbound conformation, as discussed by the authors, and so extraneous electron density was not observed for this chain.
A simple test of this hypothesis would be to perform activity assays in a buffer without HEPES, and test for competitive inhibition with increasing concentration of HEPES.