User:Wayne Decatur/SandboxRibosome
From Proteopedia
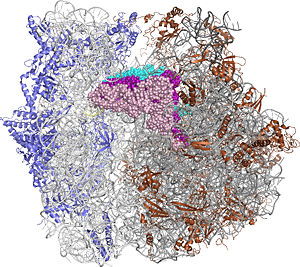
The Ribosome
The protein synthesis machine of cells
shown with the 3 transfer RNAs and messenger RNA bound.
THIS IS THE OCTOBER 15th version of the page I had made for Proteopedia Ribosome page with other parts I am working on to add to that current page
Contents |
Background
Venkatraman Ramakrishnan of the M.R.C. Laboratory of Molecular Biology in Cambridge, England; Thomas A. Steitz of Yale University; and Ada E. Yonath of the Weizmann Institute of Science in Rehovot, Israel are being awarded the the 2009 Nobel Prize in Chemistry for their for their landmark work revealing the atomic details of the molecular machine that make proteins in all cells, the ribosome. This work was the gloriously enlightening culmination of years of work, first heralded by Ada Yonath's report of crystals in 1980. Others made significant contributions to the detailed structure of this machine, as poignantly summarized by Jeremy Berg, current Director of National Institute of General Medical Sciences, in his announcement, "The Nobel committee has the daunting challenge of limiting itself to up to three laureates for each prize. Several other long-time NIGMS grantees who also contributed greatly to our understanding of the structure and function of the ribosome include Peter Moore, Harry Noller and Joachim Frank." The American Society for Biochemistry and Molecular Biology posted an announcement of the prize echoing this sentiment as well.
The ribosome is a complex composed of RNA and protein that adds up to several million daltons in size and plays a critical role in the process of decoding the genetic information stored in the genome into protein as outlined in what is now known as the Central Dogma of Molecular Biology. Specifically, the ribosome carries out the process of translation, decoding the genetic information encoded in messenger RNA, one amino acid at a time, into newly synthesized polypeptide chains. The immense size of the ribosome and each of the two individual ribosomal subunits that come together to form the complete ribosome that is active in translation made for a daunting task in structure determination. These structures were at the time, and remain, the largest asymmetric molecules solved crystallographically. In addition to providing us immense insight into the general molecular and atomic details of protein synthesis in every organism on earth, the development of new antibiotics are likely to rely on this ground-breaking work.
The small subunit of the ribosome is the main site of decoding, directing the interaction of the messenger RNA codon with the anticodon stem-loops of the proper transfer RNA. The formation of peptide bounds occurs in the large subunit where the acceptor-stems of the tRNAs are docked. However, it is important to keep in mind that in the active ribosome the two subunits are in contact via bridges and the actions in one subunit affect the other as the process of translation advances through the stages of initiation, elongation, and termination. The initial determination of the atomic resolution structures of the subunits surprisingly revealed that RNA and not protein contribute directly to forming the site of both decoding and catalysis of peptide bond synthesis, with the ribosomal proteins only acting in an ancillary role. During the elongation stage of translation, new peptides are added to the growing nascent chain that is linked to the acceptor-end of the tRNA in the peptidyl or P site, and as the nascent grows it advances into a tunnel that passes through the large subunit, called the polypeptide exit tunnel. Several factors can interact at the site of extrusion of the nascent polypeptide chain to insure proper folding or transport across a membrane. Additionally, during protein synthesis, many additional factors such as EF-Tu and EF-G interact with the ribosome to elicit decoding and peptide bond synthesis accurately and efficiently. Structures of several of these factors in complex with the ribosome, as well has intermediate states in the process, are being observed now, building upon the first atomic structures.
First Atomic-Resolution Ribosome Structures
The particular structures for which the Nobel prize was awarded were published in 2000 and were subsequently refined or improved upon. Here are the links to the Proteopedia entries:
- Yonath lab original atomic-resolution structures: Thermus thermophilus small ribosomal subunit - 1fka, improved in 1i94, 1i95, 1i96, and 1i97. Thermus thermophilus is a thermophilic eubacteria.
- Ramakrishnan lab original atomic-resolution structures: Thermus thermophilus small ribosomal subunit -1fjf which was later refined to 1j5e. Related: in complex with the antibiotics streptomycin, spectinomycin, and paromomycin in 1fjg; in complex with tetracycline in 1hnw, pactamycin in 1hnx, hygromycin B in 1hnz.
- Steitz and Moore lab original atomic-resolution structures: Haloarcula marismortui large ribosomal subunit - 1ffk and later refined to give 1jj2, and then refined to give 1s72. Related: 1ffz, 1fg0. Haloracula is a halophilic archaea.
Additional Ribosome Structures
| |||||||
·· ·· ·· ·· ·· |
Several other ribosome structures have now been published, and here are just a few of these entries in Proteopedia (with apologies to the authors of those not yet listed):
- 2j00 and 2j01 are the subunits of the 70S ribosome structure from the Ramakrishnan lab; the aminoglycoside antibiotic paromomycin is present as well.
- 1gix and 1giy are the subunits of the 70S ribosome structure determined by the Noller lab, the first for the 70S at near-atomic resolution; more of the mRNA chain is seen in 1jgo.
- 2i2u and 2i2v are the subunits of the E. coli ribosome at 3.2 Å as solved by the Cate lab. 2i2t and 2i2p are related structures.
- 2gya and 2gy9 are the subunits of a complete E. coli ribosome as determined by cryo-EM by Joachim Frank's lab.
- 1ibk is paromomycin bound to the small subunit.
- 1ibm is the small subunit with an mRNA analog bound and an anticodon stem loop bound to the A site.
- Small subunit bound to near-cognate tRNA anticodon stem-loop: 1n32, 1n33, 1n34, 1n36
- Macrolide, lincosamide, streptogramin B, and ketolide antibiotics bound to the large subunit, which impacts mechanisms of drug resistance:1yi2, 1yj2, 1yit, 1yhq, 1yjn, 1yij, 1yj9
- E. coli 70S ribosome in complex with the atypical aminoglycoside antibiotic hygromycin B: 3df1, 3df2,3df3, and 3df4.
- Ribosome Recycling Factor bound to the 70S Ribosome: 2v46, 2v47, 2v48, 2v49
- 2ow8 and 1vsa are the subunits of a 70S-tRNA-mRNA complex from the Noller lab.
- 2e5l shows the small subunit with an mRNA mimic bound and the Shine-Dalgarno and anit-Shine-Dalgarno sequences interacting.
- Structures of the 30S bound with anticodon stem-loops from tRNAs that facilitate frame-shifting: 2uxb, 2uxc, and 2uxd
- Initation factor 1 bound to the small subunit: 1hr0
- 70S Ribosome in complex with mRNA, paromomycin, acylated A- And P-Site tRNAs, and E-Site tRNA: 2wdg, 2wdh, 2wdi, 2wdj, 2wdk, 2wdl, 2wdm, and 2wdn
- E. coli 70S ribosome intermediates in a key conformational change: 3i1m, 3i1n, 3i1o, 3i1p, 3i1q, 3i1r,3i1s,3i1t,3i1z, 3i20, 3i21, and 3i22.
- E. coli ribosome in complex with the atypical aminoglycoside antibiotic hygromycin B: 3df1, 3df2,3df3, and 3df4.
- Structural basis of a mechanism of hydrolytic release of the newly formed polypeptide by the large subunit that may be analogous to that used by release factors:3cma, 3cme
- Elongation factor P bound to the 70S ribosome: 3huw, 3hux, 3huy, 3huz.
Scenes introducing the Ribosome
Three tRNAs and an mRNA are bound.
The anticodon stem-loops of A-site tRNA, P-site tRNA, and E-site tRNA stick into the 30S subunit of the ribosome.
The anticodon stem-loops of the A-site tRNA and P-site tRNA hydrogen bond to the messenger RNA.