User:Marcin Jozef Suskiewicz/Sandbox Parvin/
From Proteopedia
Alpha-parvin[1], also known as actopaxin[2] or CH domain-containing integrin-linked kinase (ILK)-binding protein (CH-ILK-BP)[3] is an adapter protein known to interact with a number of focal adhesion proteins leading to focal adhesion stabilisation. Knock-out analysis confirmed it to be essential for efficient directional cell migration during embryogenesis in mice[4]. Spatially and temporarily regulated dynamic changes in the phosphorylation status of alpha-parvin at serines 4 and 8 and consequent changes in affinities towards its binding partners (icluding CdGAP, TESK1 and possibly others, e.g. ILK) may be responsible for 1) focal adhesion turnover (disassembly of old adhesions, assembly of new ones) and 2) actin cytoskeleton reorganization, two interrelated processes contributing to cell migration.[5][6][7][8]
Contents |
Biological significance
Focal adhesions and cell migration
The extracellular matrix (ECM) composed of matrix molecules such as collagens, laminins, proteoglycans and non-matrix proteins including growth factors, provides physical support and mediates bidirectional flow of signals in all tissues and organs[9]. Cell-ECM attachments, critical for appropriate localization and functioning of cells, are mediated by integrins.[10]The engagement of integrins to the ECM results in the recruitment of various proteins to the cell periphery and consequent assembly of dot-like focal complexes (FXs or nascent adhesions), which can mature to much larger focal adhesions (FAs or focal contacts). Focal adhesions are bound to the integrins on one hand and to the actin cytoskeleton on the other, thus accounting for the mechanical continuity and signal transduction between the ECM outside the cell and the cytoskeleton inside the cell.[11] The complex FA network (the adhesome), is composed of, among others, actin regulators, Tyr and Ser/Thr kinases and phosphatases, phosphatidylinositols and their modifiers, Rho family GTPases and their effectors. Interactions between these components are mediated by various adapter proteins, including talin, vinculin, paxillin and alpha-parvin[12]. FAs can grow and shrink and, even in stationary cells, disappear on average tens of minutes after maturation.[13] The spatially and temporarily regulated FA turnover together with changes in cytoskeletal contractability enable cell migration in development and regeneration.[14][15]
Alpha-parvin's interactions in the adhesome
Alpha-parvin is thus a component of the adhesome, but a more basic unit of which it is part is the IPP (ILK-PINCH-parvin) complex. The central component of this complex, ILK, binds both PINCH and alpha-parvin, which do not interact directly with each other, and it is by means of ILK's pseudokinase domain that the complex is recruited to the cytoplasmic tail of integrins. The IPP complex is important for the correct FA assembly and promotes anti-apoptotic signalling[16]. Alpha-parvin is also known to interact with a number of other proteins, including F-actin[17], paxillin[18][19], paxillin's homologue HIC5[20], TESK1[21], CdGAP[22] and alphaPIX[23].
Alpha-parvin phosphorylation and cell migration
Alpha-parvin possesses 6 putative proline-directed serine/threonine phosphorylation targets (residues 4, 8, 14, 16, 19, 61), of which serines 4 and 8 were shown to be the most important. Phosphorylation of alpha-parvin at serines 4 and 8 is correlated with the tightly regulated process of FA turnover during cell migration. Firstly, phosphorylation of these residues by cyclin B1/cdc2 is observed in the context of mitosis, whereby it contributes to FA disassembly required for cell-rounding prior to cell division, suggesting it may cause a similar effect during cell migration.[24] Indeed, in migrating cells these residues are observed to be phosphorylated[25], likely as a result of MAP kinase[26] and/or PI3K[27]. Secondly, phosphomimetic mutations of serines 4 and 8 to aspartates result in faster migration and spreading, while mutations preventing phosphorylation impair these processes.[28] Finally, as has already been mentioned in the introduction, knock-out mice phenotype (embryonic lethality due to severe cardiovascular defects) suggests alpha-parvin deficiency results in impaired directional migration of endothelial cells during embryonic development, heart development in particular.[29] The macroscopic effects of alpha-parvin phosphorylation likely result from the altered affinity for its binding partners. So far it has been demonstrated that phosphorylation at serines 4 and 8 affects binding of alpha-parvin to TESK1 and CdGAP. When TESK1 is bound to alpha-parvin it is prevented from severing actin fibres. TESK1 is thought to be released from inhibition upon alpha-parvin phosphorylation and this can contribute to the decomposition of actin fibres and FA disassembly.[30] CdGAP, on the other hand, is involved in the regulation of small GTPase signalling, which accounts for changes in cytoskeletal contractability during cell migration.[31] Evidence suggesting that also the interaction with ILK is affected by phosphorylation is not strong[32][33], but it is likely that this or yet other binding partners bind in the phosphorylation-dependent manner.
Structure and function
Domain composition
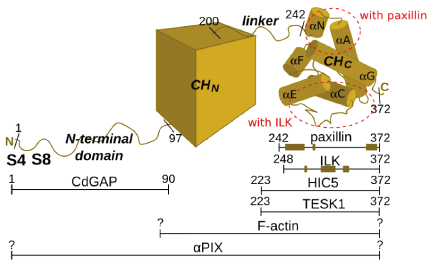
Alpha-parvin's structure can be divided into four regions: 1) N-terminal flexible domain (residues 1-96), 2) N-terminal CH domain (97-200 according to SMART[34]), 3) linker region (201-241) and 4) C-terminal CH domain (242-372 identified by limited subtilisin proteolysis[35]). Most interactions of alpha-parvin are mapped to the C-terminal CH domain, but CdGAP and perhaps alphaPIX or other, yet unknown partners, interact with the N-terminal flexibile domain. This flexible domain seems to lack a well defined 3D structure and can therefore be classified as a putative intrinsically disordered region. The interactions of this regions with the binding partners are therefore likely to be characterized by relatively low affinity, but high affinity nonetheless.[36] The abovementioned phosphorylation sites (serines 4 and 8) involved in focal adhesion regulation are located in this segment, which makes them so called disorder-enhanced phosphorylation sites.[37][38]
C-terminal CH domain
The calponin-homology (CH) domains are helical structural units around 100 amino acids long. They comprise at least four helices, three of them forming a helical bundle. CH domains usually comprise elements of big multidomain proteins and are present either in singlet[39] or duplex/tandem arrangement.[40] The tandem arrangement of CH domains is often associated with F-actin binding[41][42] (and is thus called actin-binding domain or ABD), but generally CH domains seem to be characterized by functional plasticity and ability to bind various structural motifs.[43] In the case of alpha-parvin, the interactions of CH domains with both F-actin and other partners (paxillin, ILK) are observed. The interactions with paxillin and ILK are mediated by a single CH domain, the C-terminal one. This domain has attracted most attention. While no full-length structure of alpha-parvin has been solved to date, the structure of the C-terminal CH domain, on its own[44] and in complexes (with paxillin[45][46] and the pseudokinase domain of ILK[47]) are available.
| |||||||||||
References
- ↑ Olski TM, Noegel AA, Korenbaum E. Parvin, a 42 kDa focal adhesion protein, related to the alpha-actinin superfamily. J Cell Sci. 2001 Feb;114(Pt 3):525-38. PMID:11171322
- ↑ Nikolopoulos SN, Turner CE. Actopaxin, a new focal adhesion protein that binds paxillin LD motifs and actin and regulates cell adhesion. J Cell Biol. 2000 Dec 25;151(7):1435-48. PMID:11134073
- ↑ Tu Y, Huang Y, Zhang Y, Hua Y, Wu C. A new focal adhesion protein that interacts with integrin-linked kinase and regulates cell adhesion and spreading. J Cell Biol. 2001 Apr 30;153(3):585-98. PMID:11331308
- ↑ Montanez E, Wickstrom SA, Altstatter J, Chu H, Fassler R. Alpha-parvin controls vascular mural cell recruitment to vessel wall by regulating RhoA/ROCK signalling. EMBO J. 2009 Oct 21;28(20):3132-44. Epub 2009 Oct 1. PMID:19798050 doi:10.1038/emboj.2009.295
- ↑ Clarke DM, Brown MC, LaLonde DP, Turner CE. Phosphorylation of actopaxin regulates cell spreading and migration. J Cell Biol. 2004 Sep 13;166(6):901-12. Epub 2004 Sep 7. PMID:15353548 doi:10.1083/jcb.200404024
- ↑ LaLonde DP, Brown MC, Bouverat BP, Turner CE. Actopaxin interacts with TESK1 to regulate cell spreading on fibronectin. J Biol Chem. 2005 Jun 3;280(22):21680-8. Epub 2005 Apr 6. PMID:15817463 doi:10.1074/jbc.M500752200
- ↑ LaLonde DP, Grubinger M, Lamarche-Vane N, Turner CE. CdGAP associates with actopaxin to regulate integrin-dependent changes in cell morphology and motility. Curr Biol. 2006 Jul 25;16(14):1375-85. PMID:16860736 doi:S0960-9822(06)01676-9
- ↑ Yang Y, Guo L, Blattner SM, Mundel P, Kretzler M, Wu C. Formation and phosphorylation of the PINCH-1-integrin linked kinase-alpha-parvin complex are important for regulation of renal glomerular podocyte adhesion, architecture, and survival. J Am Soc Nephrol. 2005 Jul;16(7):1966-76. Epub 2005 May 4. PMID:15872073 doi:10.1681/ASN.2004121112
- ↑ Hynes RO. The extracellular matrix: not just pretty fibrils. Science. 2009 Nov 27;326(5957):1216-9. PMID:19965464 doi:10.1126/science.1176009
- ↑ Barczyk M, Carracedo S, Gullberg D. Integrins. Cell Tissue Res. 2010 Jan;339(1):269-80. Epub 2009 Aug 20. PMID:19693543 doi:10.1007/s00441-009-0834-6
- ↑ Wolfenson H, Henis YI, Geiger B, Bershadsky AD. The heel and toe of the cell's foot: a multifaceted approach for understanding the structure and dynamics of focal adhesions. Cell Motil Cytoskeleton. 2009 Nov;66(11):1017-29. PMID:19598236 doi:10.1002/cm.20410
- ↑ Zaidel-Bar R, Itzkovitz S, Ma'ayan A, Iyengar R, Geiger B. Functional atlas of the integrin adhesome. Nat Cell Biol. 2007 Aug;9(8):858-67. PMID:17671451 doi:10.1038/ncb0807-858
- ↑ Wolfenson H, Henis YI, Geiger B, Bershadsky AD. The heel and toe of the cell's foot: a multifaceted approach for understanding the structure and dynamics of focal adhesions. Cell Motil Cytoskeleton. 2009 Nov;66(11):1017-29. PMID:19598236 doi:10.1002/cm.20410
- ↑ Zaidel-Bar R, Geiger B. The switchable integrin adhesome. J Cell Sci. 2010 May 1;123(Pt 9):1385-8. PMID:20410370 doi:10.1242/jcs.066183
- ↑ Gardel ML, Schneider IC, Aratyn-Schaus Y, Waterman CM. Mechanical integration of actin and adhesion dynamics in cell migration. Annu Rev Cell Dev Biol. 2010 Nov 10;26:315-33. PMID:19575647 doi:10.1146/annurev.cellbio.011209.122036
- ↑ Attwell S, Mills J, Troussard A, Wu C, Dedhar S. Integration of cell attachment, cytoskeletal localization, and signaling by integrin-linked kinase (ILK), CH-ILKBP, and the tumor suppressor PTEN. Mol Biol Cell. 2003 Dec;14(12):4813-25. Epub 2003 Sep 5. PMID:12960424 doi:http://dx.doi.org/10.1091/mbc.E03-05-0308
- ↑ Olski TM, Noegel AA, Korenbaum E. Parvin, a 42 kDa focal adhesion protein, related to the alpha-actinin superfamily. J Cell Sci. 2001 Feb;114(Pt 3):525-38. PMID:11171322
- ↑ Wang X, Fukuda K, Byeon IJ, Velyvis A, Wu C, Gronenborn A, Qin J. The structure of alpha-parvin CH2-paxillin LD1 complex reveals a novel modular recognition for focal adhesion assembly. J Biol Chem. 2008 Jul 25;283(30):21113-9. Epub 2008 May 28. PMID:18508764 doi:10.1074/jbc.M801270200
- ↑ Lorenz S, Vakonakis I, Lowe ED, Campbell ID, Noble ME, Hoellerer MK. Structural analysis of the interactions between paxillin LD motifs and alpha-parvin. Structure. 2008 Oct 8;16(10):1521-31. PMID:18940607 doi:10.1016/j.str.2008.08.007
- ↑ Nikolopoulos SN, Turner CE. Actopaxin, a new focal adhesion protein that binds paxillin LD motifs and actin and regulates cell adhesion. J Cell Biol. 2000 Dec 25;151(7):1435-48. PMID:11134073
- ↑ LaLonde DP, Brown MC, Bouverat BP, Turner CE. Actopaxin interacts with TESK1 to regulate cell spreading on fibronectin. J Biol Chem. 2005 Jun 3;280(22):21680-8. Epub 2005 Apr 6. PMID:15817463 doi:10.1074/jbc.M500752200
- ↑ LaLonde DP, Grubinger M, Lamarche-Vane N, Turner CE. CdGAP associates with actopaxin to regulate integrin-dependent changes in cell morphology and motility. Curr Biol. 2006 Jul 25;16(14):1375-85. PMID:16860736 doi:S0960-9822(06)01676-9
- ↑ LaLonde DP, Grubinger M, Lamarche-Vane N, Turner CE. CdGAP associates with actopaxin to regulate integrin-dependent changes in cell morphology and motility. Curr Biol. 2006 Jul 25;16(14):1375-85. PMID:16860736 doi:S0960-9822(06)01676-9
- ↑ Curtis M, Nikolopoulos SN, Turner CE. Actopaxin is phosphorylated during mitosis and is a substrate for cyclin B1/cdc2 kinase. Biochem J. 2002 Apr 15;363(Pt 2):233-42. PMID:11931650
- ↑ Clarke DM, Brown MC, LaLonde DP, Turner CE. Phosphorylation of actopaxin regulates cell spreading and migration. J Cell Biol. 2004 Sep 13;166(6):901-12. Epub 2004 Sep 7. PMID:15353548 doi:10.1083/jcb.200404024
- ↑ Ishibe S, Joly D, Zhu X, Cantley LG. Phosphorylation-dependent paxillin-ERK association mediates hepatocyte growth factor-stimulated epithelial morphogenesis. Mol Cell. 2003 Nov;12(5):1275-85. PMID:14636584
- ↑ Attwell S, Mills J, Troussard A, Wu C, Dedhar S. Integration of cell attachment, cytoskeletal localization, and signaling by integrin-linked kinase (ILK), CH-ILKBP, and the tumor suppressor PTEN. Mol Biol Cell. 2003 Dec;14(12):4813-25. Epub 2003 Sep 5. PMID:12960424 doi:http://dx.doi.org/10.1091/mbc.E03-05-0308
- ↑ Clarke DM, Brown MC, LaLonde DP, Turner CE. Phosphorylation of actopaxin regulates cell spreading and migration. J Cell Biol. 2004 Sep 13;166(6):901-12. Epub 2004 Sep 7. PMID:15353548 doi:10.1083/jcb.200404024
- ↑ Montanez E, Wickstrom SA, Altstatter J, Chu H, Fassler R. Alpha-parvin controls vascular mural cell recruitment to vessel wall by regulating RhoA/ROCK signalling. EMBO J. 2009 Oct 21;28(20):3132-44. Epub 2009 Oct 1. PMID:19798050 doi:10.1038/emboj.2009.295
- ↑ LaLonde DP, Brown MC, Bouverat BP, Turner CE. Actopaxin interacts with TESK1 to regulate cell spreading on fibronectin. J Biol Chem. 2005 Jun 3;280(22):21680-8. Epub 2005 Apr 6. PMID:15817463 doi:10.1074/jbc.M500752200
- ↑ LaLonde DP, Grubinger M, Lamarche-Vane N, Turner CE. CdGAP associates with actopaxin to regulate integrin-dependent changes in cell morphology and motility. Curr Biol. 2006 Jul 25;16(14):1375-85. PMID:16860736 doi:S0960-9822(06)01676-9
- ↑ Yang Y, Guo L, Blattner SM, Mundel P, Kretzler M, Wu C. Formation and phosphorylation of the PINCH-1-integrin linked kinase-alpha-parvin complex are important for regulation of renal glomerular podocyte adhesion, architecture, and survival. J Am Soc Nephrol. 2005 Jul;16(7):1966-76. Epub 2005 May 4. PMID:15872073 doi:10.1681/ASN.2004121112
- ↑ Attwell S, Mills J, Troussard A, Wu C, Dedhar S. Integration of cell attachment, cytoskeletal localization, and signaling by integrin-linked kinase (ILK), CH-ILKBP, and the tumor suppressor PTEN. Mol Biol Cell. 2003 Dec;14(12):4813-25. Epub 2003 Sep 5. PMID:12960424 doi:http://dx.doi.org/10.1091/mbc.E03-05-0308
- ↑ Schultz J, Milpetz F, Bork P, Ponting CP. SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci U S A. 1998 May 26;95(11):5857-64. PMID:9600884
- ↑ Lorenz S, Vakonakis I, Lowe ED, Campbell ID, Noble ME, Hoellerer MK. Structural analysis of the interactions between paxillin LD motifs and alpha-parvin. Structure. 2008 Oct 8;16(10):1521-31. PMID:18940607 doi:10.1016/j.str.2008.08.007
- ↑ Espinoza-Fonseca LM. Reconciling binding mechanisms of intrinsically disordered proteins. Biochem Biophys Res Commun. 2009 May 8;382(3):479-82. Epub 2009 Mar 3. PMID:19265676 doi:10.1016/j.bbrc.2009.02.151
- ↑ Iakoucheva LM, Radivojac P, Brown CJ, O'Connor TR, Sikes JG, Obradovic Z, Dunker AK. The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Res. 2004 Feb 11;32(3):1037-49. Print 2004. PMID:14960716 doi:10.1093/nar/gkh253
- ↑ Collins MO, Yu L, Campuzano I, Grant SG, Choudhary JS. Phosphoproteomic analysis of the mouse brain cytosol reveals a predominance of protein phosphorylation in regions of intrinsic sequence disorder. Mol Cell Proteomics. 2008 Jul;7(7):1331-48. Epub 2008 Apr 3. PMID:18388127 doi:10.1074/mcp.M700564-MCP200
- ↑ Friedberg F. Singlet CH domain containing human multidomain proteins: an inventory. Mol Biol Rep. 2010 Mar;37(3):1531-9. Epub 2009 May 21. PMID:19459066 doi:10.1007/s11033-009-9554-y
- ↑ Friedberg F. Duplex (or quadruplet) CH domain containing human multidomain proteins: an inventory. Mol Biol Rep. 2010 Apr;37(4):1707-16. Epub 2009 Jun 30. PMID:19565353 doi:10.1007/s11033-009-9589-0
- ↑ Stradal T, Kranewitter W, Winder SJ, Gimona M. CH domains revisited. FEBS Lett. 1998 Jul 17;431(2):134-7. PMID:9708889
- ↑ Sjoblom B, Ylanne J, Djinovic-Carugo K. Novel structural insights into F-actin-binding and novel functions of calponin homology domains. Curr Opin Struct Biol. 2008 Dec;18(6):702-8. Epub 2008 Nov 13. PMID:18952167 doi:10.1016/j.sbi.2008.10.003
- ↑ Gimona M, Djinovic-Carugo K, Kranewitter WJ, Winder SJ. Functional plasticity of CH domains. FEBS Lett. 2002 Feb 20;513(1):98-106. PMID:11911887
- ↑ Lorenz S, Vakonakis I, Lowe ED, Campbell ID, Noble ME, Hoellerer MK. Structural analysis of the interactions between paxillin LD motifs and alpha-parvin. Structure. 2008 Oct 8;16(10):1521-31. PMID:18940607 doi:10.1016/j.str.2008.08.007
- ↑ Lorenz S, Vakonakis I, Lowe ED, Campbell ID, Noble ME, Hoellerer MK. Structural analysis of the interactions between paxillin LD motifs and alpha-parvin. Structure. 2008 Oct 8;16(10):1521-31. PMID:18940607 doi:10.1016/j.str.2008.08.007
- ↑ Wang X, Fukuda K, Byeon IJ, Velyvis A, Wu C, Gronenborn A, Qin J. The structure of alpha-parvin CH2-paxillin LD1 complex reveals a novel modular recognition for focal adhesion assembly. J Biol Chem. 2008 Jul 25;283(30):21113-9. Epub 2008 May 28. PMID:18508764 doi:10.1074/jbc.M801270200
- ↑ Fukuda K, Gupta S, Chen K, Wu C, Qin J. The pseudoactive site of ILK is essential for its binding to alpha-Parvin and localization to focal adhesions. Mol Cell. 2009 Dec 11;36(5):819-30. PMID:20005845 doi:10.1016/j.molcel.2009.11.028
- ↑ Lorenz S, Vakonakis I, Lowe ED, Campbell ID, Noble ME, Hoellerer MK. Structural analysis of the interactions between paxillin LD motifs and alpha-parvin. Structure. 2008 Oct 8;16(10):1521-31. PMID:18940607 doi:10.1016/j.str.2008.08.007